Probe CUST_25579_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25579_PI426222305 | JHI_St_60k_v1 | DMT400037335 | TGCGAAATGCCTGATTATGCTTTGGTTATATATTGAAGGCTTTGGACAAGATTCGCTTTG |
All Microarray Probes Designed to Gene DMG400014405
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25643_PI426222305 | JHI_St_60k_v1 | DMT400037336 | ATTTGCTCGATTTCTTCTATCGTCGTTGAAGTTTAGCAGCAGCATCGCAGATCTGTAATG |
| CUST_25579_PI426222305 | JHI_St_60k_v1 | DMT400037335 | TGCGAAATGCCTGATTATGCTTTGGTTATATATTGAAGGCTTTGGACAAGATTCGCTTTG |
Microarray Signals from CUST_25579_PI426222305
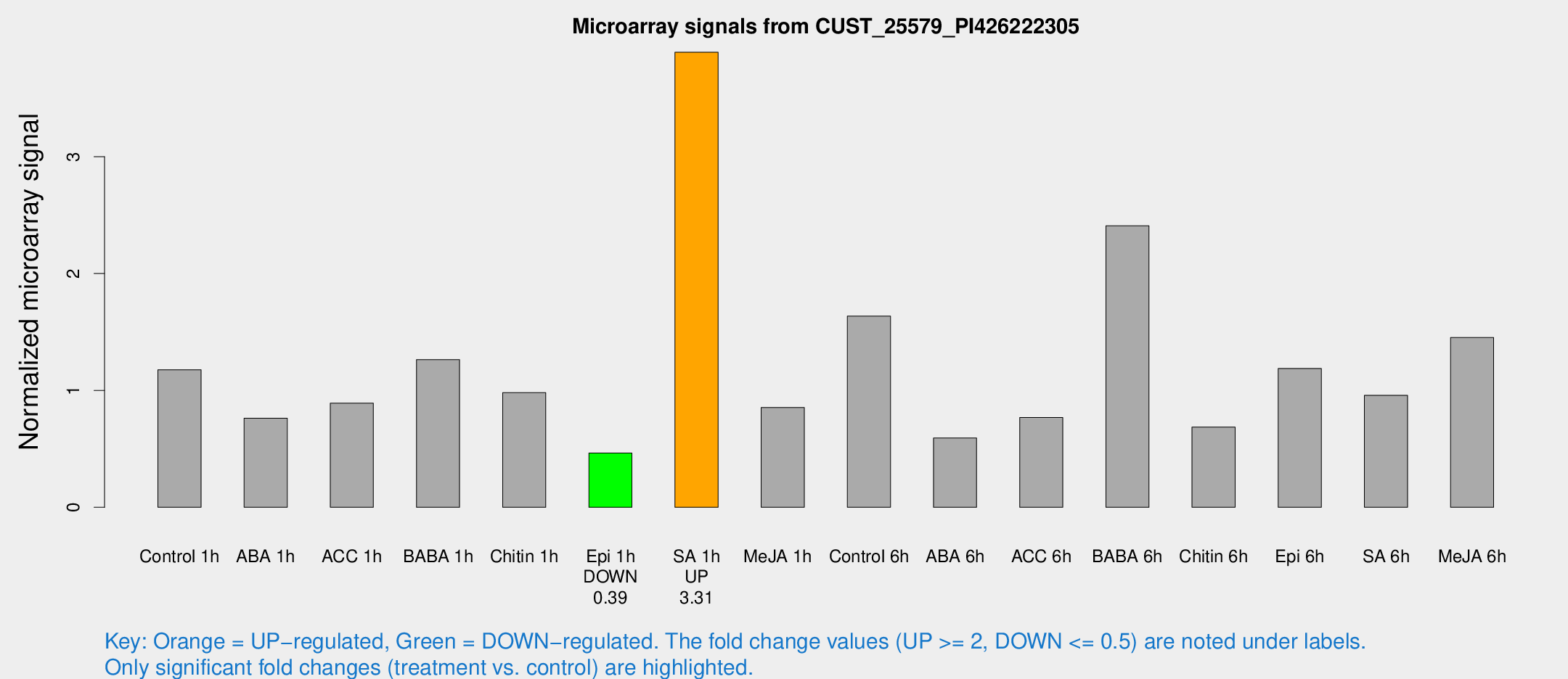
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 19.6485 | 3.82526 | 1.17692 | 0.243227 |
| ABA 1h | 13.1443 | 5.88982 | 0.762212 | 0.316217 |
| ACC 1h | 15.1203 | 4.76938 | 0.890701 | 0.318292 |
| BABA 1h | 20.1528 | 4.10677 | 1.26353 | 0.272228 |
| Chitin 1h | 14.8662 | 3.72829 | 0.980691 | 0.259719 |
| Epi 1h | 6.48483 | 3.52842 | 0.464017 | 0.2516 |
| SA 1h | 67.6622 | 15.3576 | 3.89355 | 0.844983 |
| Me-JA 1h | 11.5501 | 3.81755 | 0.854182 | 0.290297 |
| Control 6h | 33.4041 | 12.9555 | 1.6352 | 0.838084 |
| ABA 6h | 11.0546 | 4.05089 | 0.592963 | 0.254169 |
| ACC 6h | 16.0749 | 4.79008 | 0.768082 | 0.279262 |
| BABA 6h | 47.0708 | 12.4183 | 2.40794 | 0.631257 |
| Chitin 6h | 12.1656 | 4.33764 | 0.686608 | 0.265143 |
| Epi 6h | 25.7602 | 9.94343 | 1.1881 | 0.683016 |
| SA 6h | 17.9164 | 6.26933 | 0.958305 | 0.330206 |
| Me-JA 6h | 28.0717 | 12.6339 | 1.45255 | 0.558473 |
Source Transcript PGSC0003DMT400037335 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G56030.2 | +2 | 0.0 | 1041 | 616/659 (93%) | heat shock protein 81-2 | chr5:22686923-22689433 FORWARD LENGTH=728 |
