Probe CUST_25497_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25497_PI426222305 | JHI_St_60k_v1 | DMT400029363 | GGATGCTGAGGCGGTCTTCAATCATACACCAAAACATGAATGAAAAAAGAAATTTTAGAT |
All Microarray Probes Designed to Gene DMG400011282
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25497_PI426222305 | JHI_St_60k_v1 | DMT400029363 | GGATGCTGAGGCGGTCTTCAATCATACACCAAAACATGAATGAAAAAAGAAATTTTAGAT |
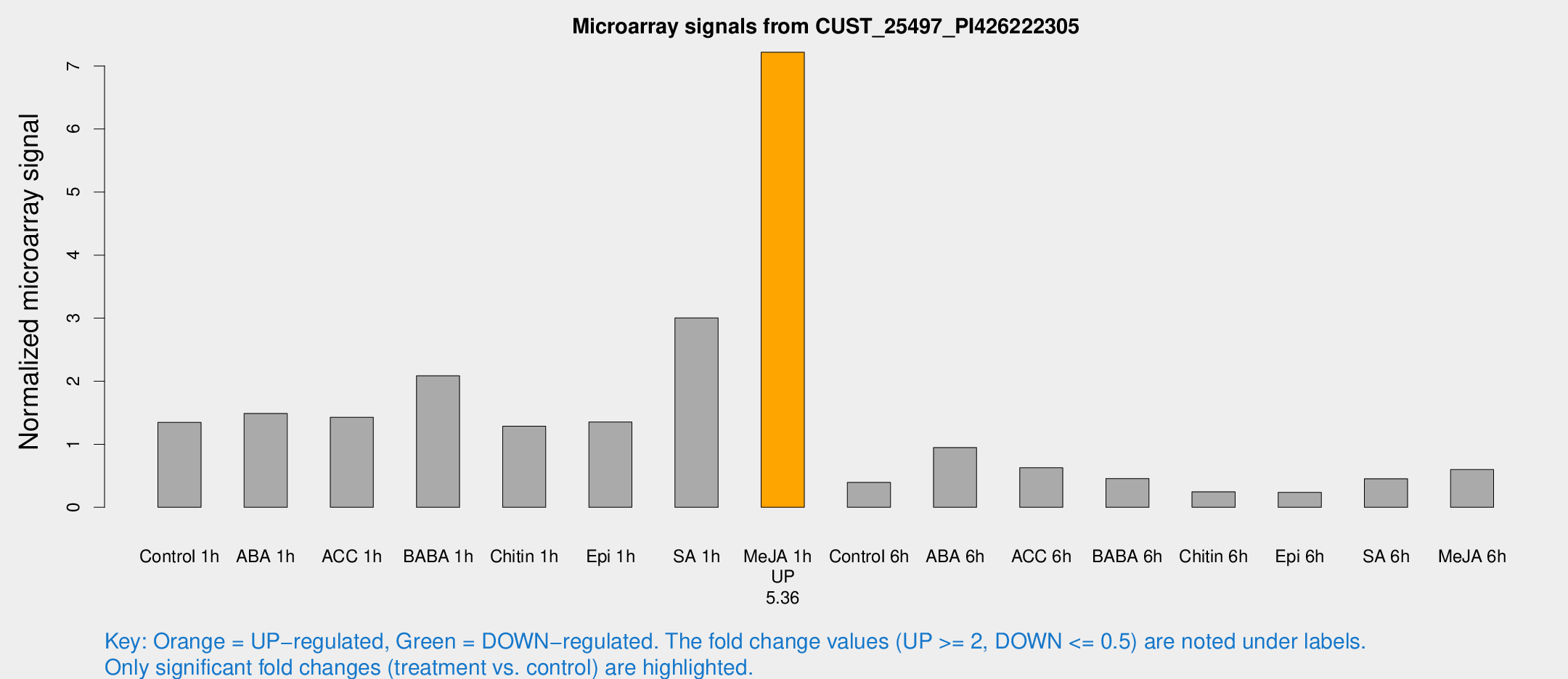
Microarray Signals from CUST_25497_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18689.3 | 3372.18 | 1.3477 | 0.19253 |
| ABA 1h | 20487 | 8033.92 | 1.48823 | 0.491411 |
| ACC 1h | 25806.5 | 13730.6 | 1.4277 | 0.787795 |
| BABA 1h | 27635.2 | 4147.64 | 2.08535 | 0.171304 |
| Chitin 1h | 15680.1 | 1440.57 | 1.28595 | 0.0742451 |
| Epi 1h | 16051.8 | 2183.06 | 1.35297 | 0.185953 |
| SA 1h | 49939 | 21629.5 | 3.0046 | 1.58879 |
| Me-JA 1h | 80777 | 11764.1 | 7.21749 | 0.477 |
| Control 6h | 5799.57 | 1746.13 | 0.394917 | 0.0878368 |
| ABA 6h | 15393.7 | 5014.08 | 0.948086 | 0.328917 |
| ACC 6h | 11520.2 | 4895.94 | 0.626548 | 0.359357 |
| BABA 6h | 7473.59 | 2198.2 | 0.455505 | 0.136086 |
| Chitin 6h | 3556.53 | 368.348 | 0.24575 | 0.0349098 |
| Epi 6h | 3619.16 | 417.366 | 0.235839 | 0.0331328 |
| SA 6h | 6098.99 | 674.683 | 0.453155 | 0.0261646 |
| Me-JA 6h | 8062.77 | 654.303 | 0.598242 | 0.0794528 |
Source Transcript PGSC0003DMT400029363 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G28237.1 | +2 | 0.0 | 565 | 275/387 (71%) | Pyridoxal-5-phosphate-dependent enzyme family protein | chr5:10207477-10213542 REVERSE LENGTH=465 |
