Probe CUST_25463_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25463_PI426222305 | JHI_St_60k_v1 | DMT400043710 | CAATCTGTAGATGAATGTTCGCGAAGCAAGAAAACAATGCCAAAATGGATGGATGAAAGA |
All Microarray Probes Designed to Gene DMG400016977
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25463_PI426222305 | JHI_St_60k_v1 | DMT400043710 | CAATCTGTAGATGAATGTTCGCGAAGCAAGAAAACAATGCCAAAATGGATGGATGAAAGA |
| CUST_25415_PI426222305 | JHI_St_60k_v1 | DMT400043708 | AAAGGATTGTTATTCCCAAAGGGAGGTTGGGAAGAAGATGAAACGATTGAGGATGCAGCG |
Microarray Signals from CUST_25463_PI426222305
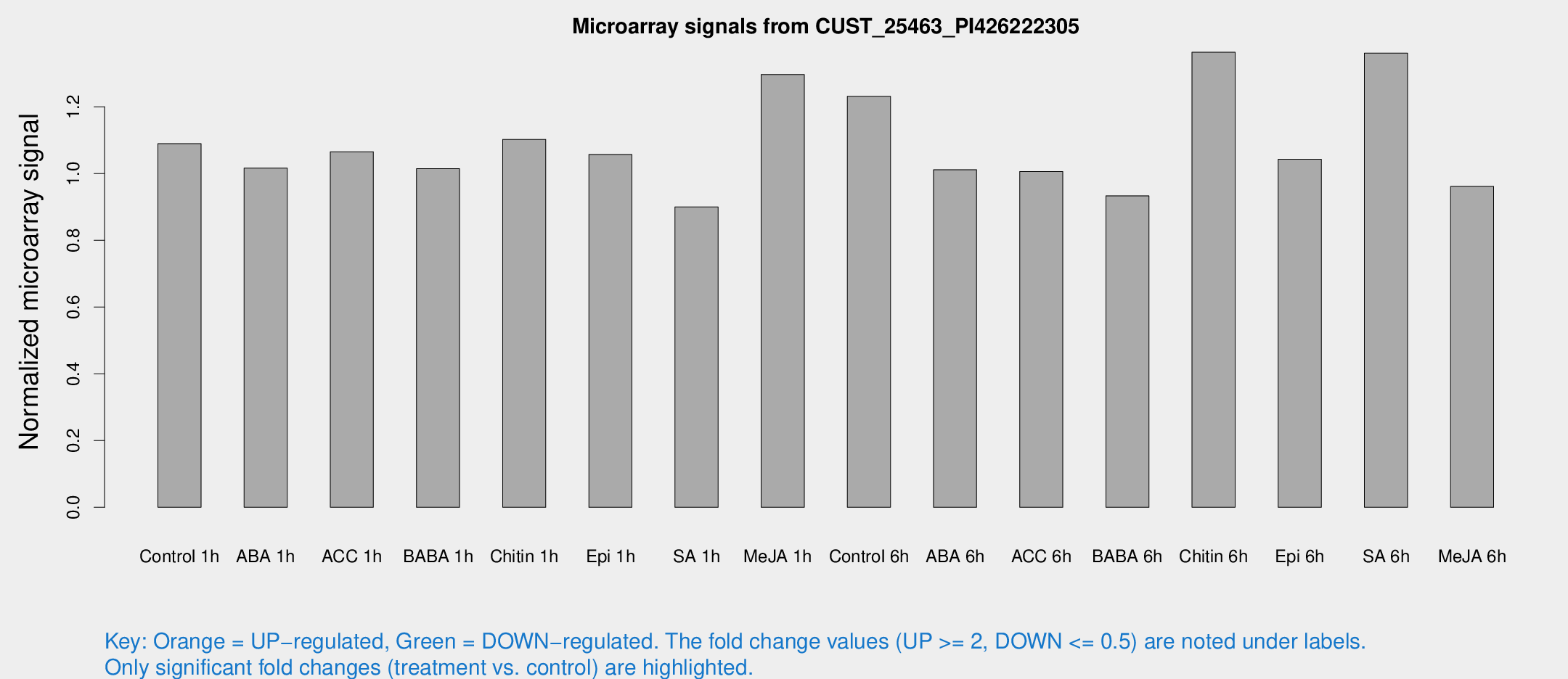
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.57763 | 3.64485 | 1.08946 | 0.546642 |
| ABA 1h | 6.17466 | 3.58202 | 1.01636 | 0.588783 |
| ACC 1h | 7.62416 | 4.46502 | 1.06531 | 0.61175 |
| BABA 1h | 6.70731 | 3.88757 | 1.01435 | 0.587758 |
| Chitin 1h | 6.84156 | 3.7476 | 1.10232 | 0.606045 |
| Epi 1h | 6.29495 | 3.6583 | 1.05708 | 0.612348 |
| SA 1h | 6.33949 | 3.64377 | 0.899727 | 0.517104 |
| Me-JA 1h | 7.32769 | 3.71051 | 1.29656 | 0.670779 |
| Control 6h | 8.65927 | 3.74574 | 1.23116 | 0.563657 |
| ABA 6h | 7.42844 | 3.97436 | 1.01139 | 0.54614 |
| ACC 6h | 8.08349 | 4.78677 | 1.00605 | 0.582649 |
| BABA 6h | 7.19491 | 4.18646 | 0.933109 | 0.540625 |
| Chitin 6h | 11.0259 | 4.26056 | 1.36342 | 0.635212 |
| Epi 6h | 8.1836 | 4.82035 | 1.04279 | 0.605002 |
| SA 6h | 9.48766 | 4.13781 | 1.36045 | 0.623747 |
| Me-JA 6h | 6.56957 | 3.8084 | 0.961306 | 0.556899 |
Source Transcript PGSC0003DMT400043710 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G01670.1 | +2 | 3e-18 | 80 | 41/101 (41%) | nudix hydrolase homolog 17 | chr2:296889-297818 REVERSE LENGTH=182 |
