Probe CUST_25446_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25446_PI426222305 | JHI_St_60k_v1 | DMT400043757 | CTGAAATCGGATGTGAATTATGGTTTCGTGCTCGACATTGGCAATACATTGAACAAGAAA |
All Microarray Probes Designed to Gene DMG400016984
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25446_PI426222305 | JHI_St_60k_v1 | DMT400043757 | CTGAAATCGGATGTGAATTATGGTTTCGTGCTCGACATTGGCAATACATTGAACAAGAAA |
Microarray Signals from CUST_25446_PI426222305
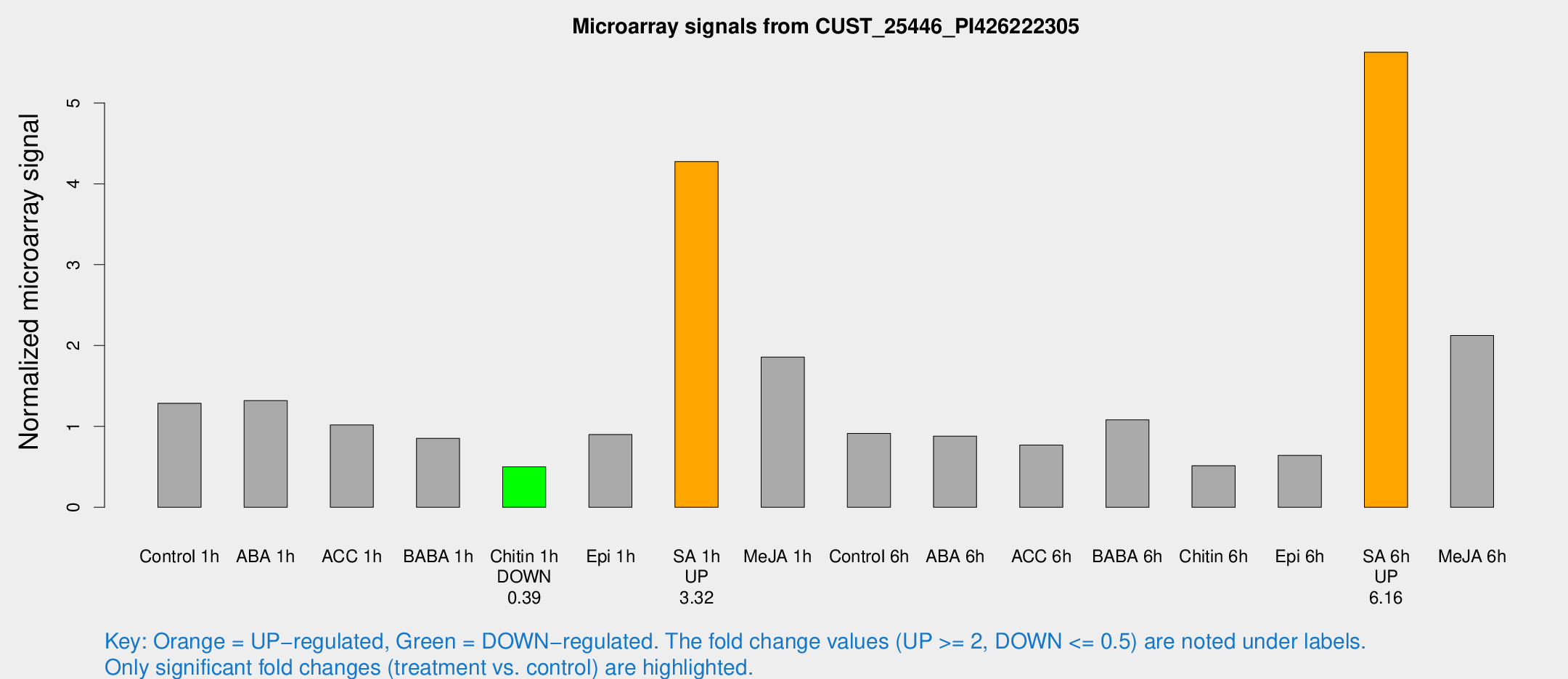
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 989.202 | 175.689 | 1.28646 | 0.139573 |
| ABA 1h | 913.422 | 194.456 | 1.31984 | 0.264982 |
| ACC 1h | 838.468 | 221.769 | 1.01857 | 0.222595 |
| BABA 1h | 654.529 | 146.407 | 0.853708 | 0.137634 |
| Chitin 1h | 346.228 | 55.7346 | 0.500579 | 0.063338 |
| Epi 1h | 581.492 | 33.6954 | 0.89952 | 0.052116 |
| SA 1h | 3444.06 | 697.318 | 4.27548 | 0.808808 |
| Me-JA 1h | 1222.97 | 350.51 | 1.85777 | 0.359321 |
| Control 6h | 745.741 | 218.317 | 0.913043 | 0.200384 |
| ABA 6h | 740.04 | 172.884 | 0.880586 | 0.167652 |
| ACC 6h | 666.787 | 73.8641 | 0.76906 | 0.157243 |
| BABA 6h | 925.686 | 144.497 | 1.08112 | 0.206583 |
| Chitin 6h | 408.878 | 23.8817 | 0.513157 | 0.0357903 |
| Epi 6h | 545.503 | 53.7214 | 0.641251 | 0.115852 |
| SA 6h | 4232.41 | 559.402 | 5.62699 | 1.11339 |
| Me-JA 6h | 1713.53 | 499.444 | 2.12445 | 0.814999 |
Source Transcript PGSC0003DMT400043757 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37980.1 | +1 | 6e-50 | 164 | 78/113 (69%) | elicitor-activated gene 3-1 | chr4:17852670-17854302 FORWARD LENGTH=357 |
