Probe CUST_25307_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25307_PI426222305 | JHI_St_60k_v1 | DMT400043759 | GGATTCGAAAGCTTCTTTCATCCATCTGCAATTTATCTATTTGTCAGGAAATGGAAAATG |
All Microarray Probes Designed to Gene DMG400016985
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25307_PI426222305 | JHI_St_60k_v1 | DMT400043759 | GGATTCGAAAGCTTCTTTCATCCATCTGCAATTTATCTATTTGTCAGGAAATGGAAAATG |
Microarray Signals from CUST_25307_PI426222305
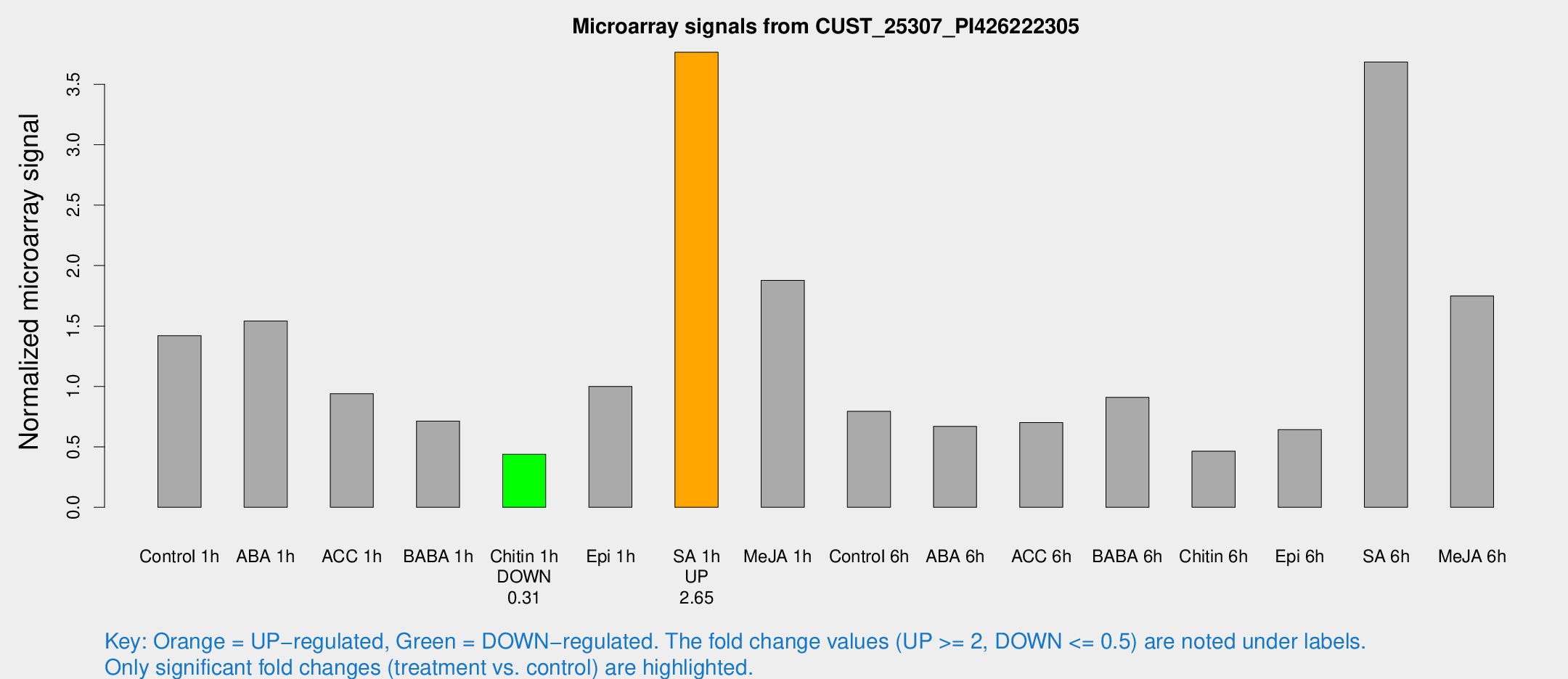
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 299.173 | 39.6698 | 1.41921 | 0.100978 |
| ABA 1h | 294.482 | 57.052 | 1.54117 | 0.275417 |
| ACC 1h | 215.804 | 58.0057 | 0.94041 | 0.211794 |
| BABA 1h | 153.843 | 37.5354 | 0.713582 | 0.136045 |
| Chitin 1h | 84.2326 | 13.6584 | 0.439083 | 0.0598388 |
| Epi 1h | 181.165 | 16.6912 | 1.00022 | 0.124524 |
| SA 1h | 841.382 | 175.882 | 3.76504 | 0.732019 |
| Me-JA 1h | 349.54 | 101.564 | 1.87801 | 0.451138 |
| Control 6h | 190.615 | 69.1715 | 0.794038 | 0.254529 |
| ABA 6h | 162.434 | 45.153 | 0.670162 | 0.191954 |
| ACC 6h | 169.385 | 21.2836 | 0.701173 | 0.152193 |
| BABA 6h | 225.122 | 58.7309 | 0.909782 | 0.273399 |
| Chitin 6h | 103.292 | 8.70523 | 0.464708 | 0.0461797 |
| Epi 6h | 154.326 | 24.2728 | 0.642883 | 0.143694 |
| SA 6h | 778.657 | 138.355 | 3.68367 | 0.717456 |
| Me-JA 6h | 392.37 | 110.774 | 1.74898 | 0.670633 |
Source Transcript PGSC0003DMT400043759 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37980.1 | +1 | 3e-17 | 75 | 34/48 (71%) | elicitor-activated gene 3-1 | chr4:17852670-17854302 FORWARD LENGTH=357 |
