Probe CUST_25220_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25220_PI426222305 | JHI_St_60k_v1 | DMT400015017 | GGTCTCTATTTGGATCAAACAATAACATGAATACGACTTCTCATCTACCCGTCAATCAAG |
All Microarray Probes Designed to Gene DMG400005860
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25220_PI426222305 | JHI_St_60k_v1 | DMT400015017 | GGTCTCTATTTGGATCAAACAATAACATGAATACGACTTCTCATCTACCCGTCAATCAAG |
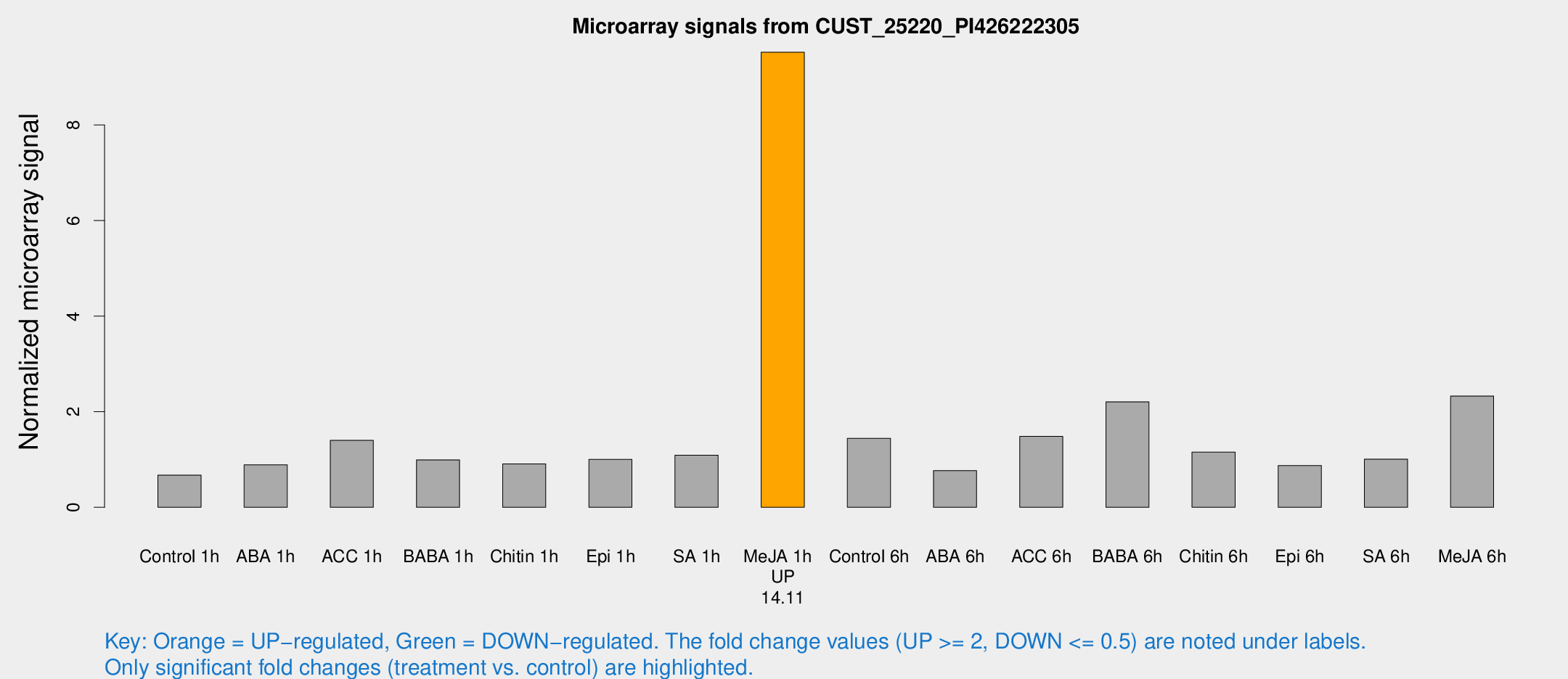
Microarray Signals from CUST_25220_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.96688 | 3.4707 | 0.674846 | 0.391126 |
| ABA 1h | 7.23738 | 3.32939 | 0.887461 | 0.443437 |
| ACC 1h | 14.1279 | 4.91666 | 1.39987 | 0.567553 |
| BABA 1h | 8.86763 | 3.79479 | 0.993028 | 0.474629 |
| Chitin 1h | 7.22445 | 3.47599 | 0.908982 | 0.437967 |
| Epi 1h | 7.90601 | 3.3841 | 1.00223 | 0.468541 |
| SA 1h | 10.7328 | 3.49581 | 1.09198 | 0.427622 |
| Me-JA 1h | 71.7042 | 15.7766 | 9.52328 | 2.00157 |
| Control 6h | 18.6066 | 11.5472 | 1.44422 | 1.04295 |
| ABA 6h | 7.26421 | 3.78129 | 0.769229 | 0.40732 |
| ACC 6h | 17.6048 | 5.87838 | 1.48416 | 0.736984 |
| BABA 6h | 35.2736 | 24.1619 | 2.20699 | 1.99522 |
| Chitin 6h | 11.2865 | 4.0731 | 1.15382 | 0.451015 |
| Epi 6h | 9.78038 | 4.23595 | 0.873523 | 0.432226 |
| SA 6h | 9.6694 | 3.8757 | 1.00751 | 0.482556 |
| Me-JA 6h | 21.9967 | 5.27083 | 2.32932 | 0.935256 |
Source Transcript PGSC0003DMT400015017 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G46760.1 | +1 | 1e-74 | 248 | 179/552 (32%) | Basic helix-loop-helix (bHLH) DNA-binding family protein | chr5:18974231-18976009 FORWARD LENGTH=592 |
