Probe CUST_25205_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25205_PI426222305 | JHI_St_60k_v1 | DMT400014932 | CCATTTCATCCAACACAAAATCCTCATGGTGTTATTCAGATGGGTTTGGCTGAAAATCAG |
All Microarray Probes Designed to Gene DMG400005825
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25205_PI426222305 | JHI_St_60k_v1 | DMT400014932 | CCATTTCATCCAACACAAAATCCTCATGGTGTTATTCAGATGGGTTTGGCTGAAAATCAG |
Microarray Signals from CUST_25205_PI426222305
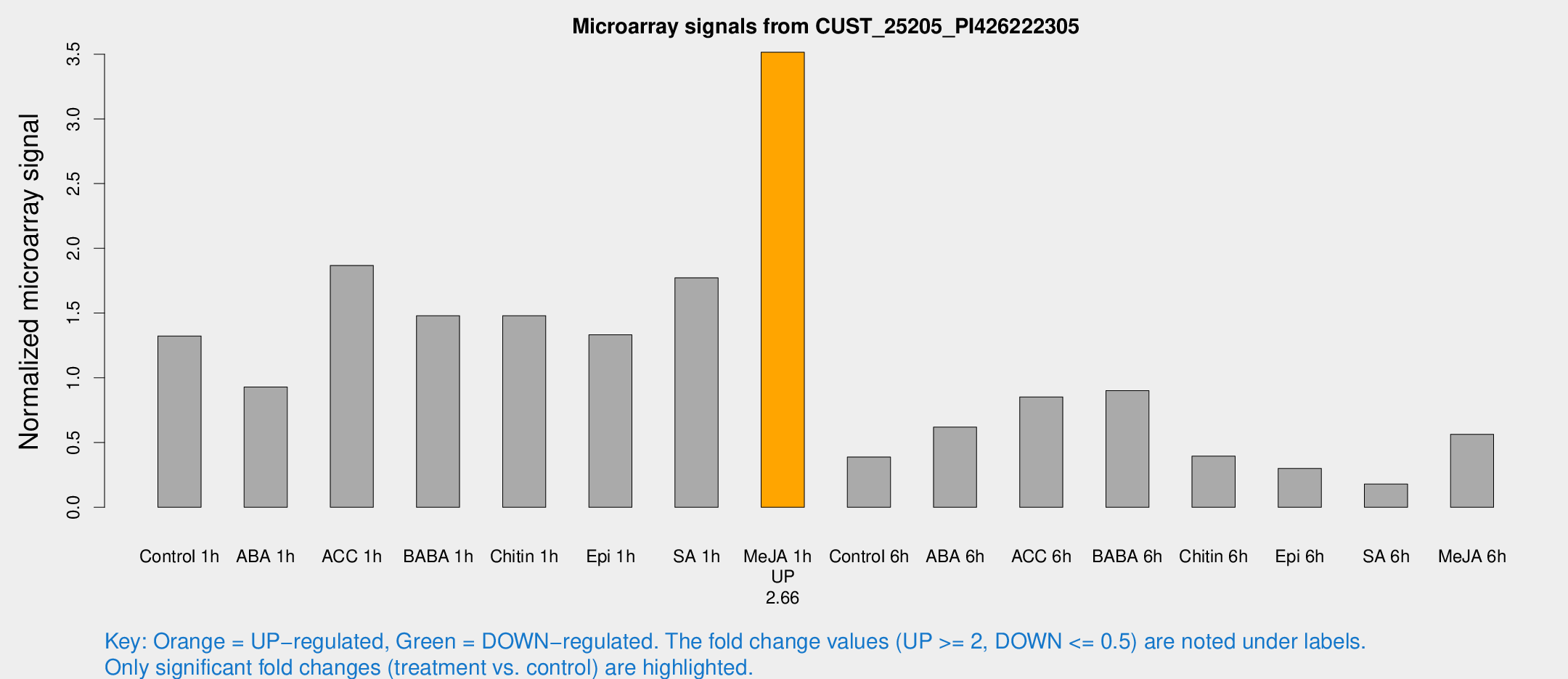
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 449.331 | 26.105 | 1.32337 | 0.076855 |
| ABA 1h | 291.986 | 57.0368 | 0.928665 | 0.163103 |
| ACC 1h | 742.783 | 269.151 | 1.86825 | 0.619624 |
| BABA 1h | 499.45 | 83.6002 | 1.47975 | 0.121377 |
| Chitin 1h | 452.833 | 26.3592 | 1.48025 | 0.147523 |
| Epi 1h | 391.268 | 22.764 | 1.33189 | 0.0774894 |
| SA 1h | 623.055 | 54.9216 | 1.77244 | 0.10267 |
| Me-JA 1h | 982.223 | 87.8649 | 3.51476 | 0.203196 |
| Control 6h | 147.485 | 41.6872 | 0.387928 | 0.107711 |
| ABA 6h | 260.249 | 83.3537 | 0.619347 | 0.262134 |
| ACC 6h | 338.244 | 46.9757 | 0.852334 | 0.0677515 |
| BABA 6h | 386.739 | 140.591 | 0.901228 | 0.295024 |
| Chitin 6h | 154.892 | 38.6815 | 0.395151 | 0.129897 |
| Epi 6h | 115.716 | 10.6789 | 0.299617 | 0.0513272 |
| SA 6h | 71.1687 | 24.8999 | 0.179247 | 0.106829 |
| Me-JA 6h | 200.562 | 46.0103 | 0.564083 | 0.0815635 |
Source Transcript PGSC0003DMT400014932 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G11280.1 | +2 | 0.0 | 633 | 308/464 (66%) | 1-aminocyclopropane-1-carboxylic acid (acc) synthase 6 | chr4:6864168-6865922 FORWARD LENGTH=495 |
