Probe CUST_25007_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25007_PI426222305 | JHI_St_60k_v1 | DMT400084732 | ATAAATTGCTTCAGCTTCCTTCTGCTTGCAAGCTCACGAACGCTAGTATTTCTAACTGTC |
All Microarray Probes Designed to Gene DMG400034309
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25007_PI426222305 | JHI_St_60k_v1 | DMT400084732 | ATAAATTGCTTCAGCTTCCTTCTGCTTGCAAGCTCACGAACGCTAGTATTTCTAACTGTC |
| CUST_24997_PI426222305 | JHI_St_60k_v1 | DMT400084731 | GTAACTATGCATCGGCCTTTGCATAAATATCGTGTTCAGTTAAACTCCTTCATATGAATT |
Microarray Signals from CUST_25007_PI426222305
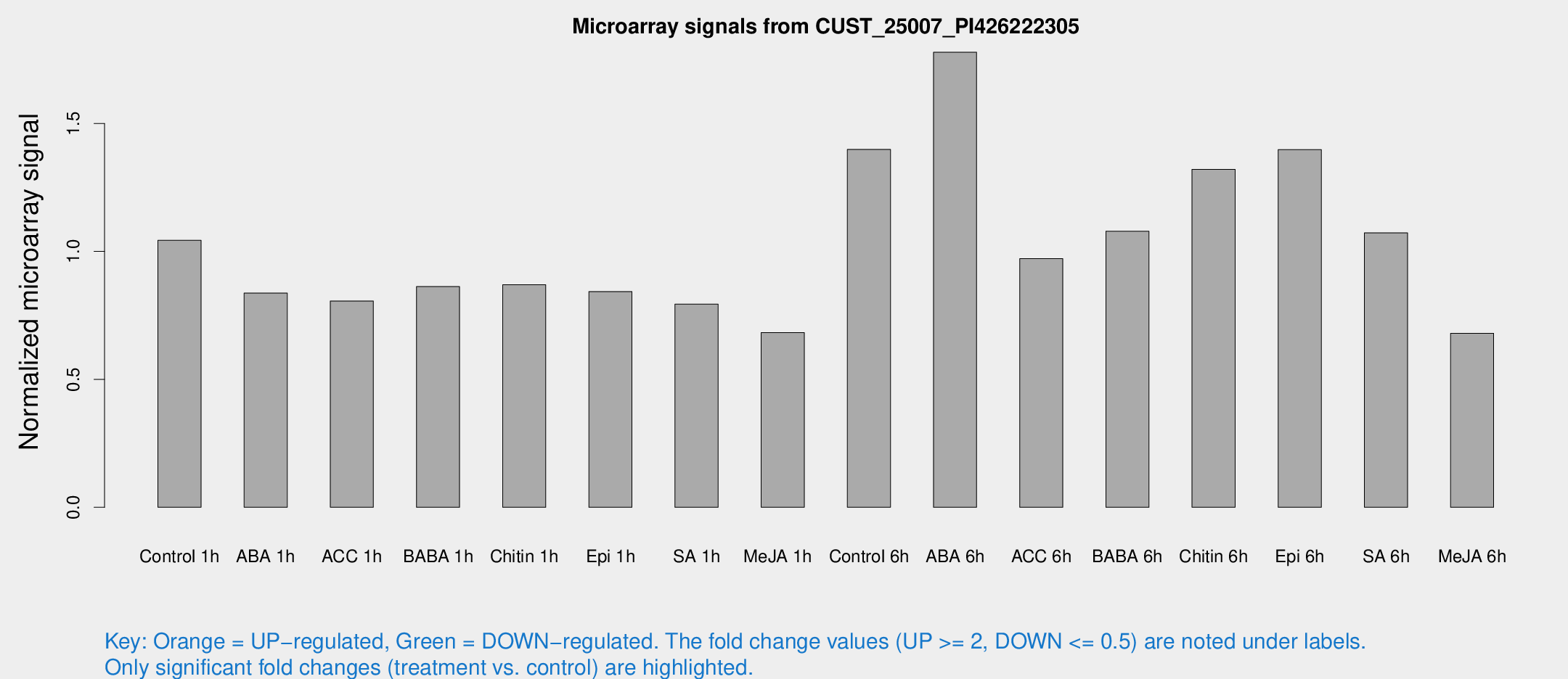
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3697.1 | 349.88 | 1.04383 | 0.060275 |
| ABA 1h | 2607.69 | 151.079 | 0.837069 | 0.0483418 |
| ACC 1h | 3193.31 | 930.316 | 0.806408 | 0.201511 |
| BABA 1h | 2991.56 | 463.815 | 0.862469 | 0.0680378 |
| Chitin 1h | 2762.34 | 212.047 | 0.869881 | 0.0696439 |
| Epi 1h | 2565.49 | 148.351 | 0.843192 | 0.0525825 |
| SA 1h | 2901.84 | 323.09 | 0.794204 | 0.0689041 |
| Me-JA 1h | 1962.44 | 113.686 | 0.682812 | 0.0770551 |
| Control 6h | 5228.69 | 1163.21 | 1.39886 | 0.23987 |
| ABA 6h | 6678.24 | 518.267 | 1.77854 | 0.10269 |
| ACC 6h | 4064.12 | 806.696 | 0.97164 | 0.0946636 |
| BABA 6h | 4340.67 | 642.93 | 1.07898 | 0.137639 |
| Chitin 6h | 4948.21 | 286.47 | 1.32065 | 0.076257 |
| Epi 6h | 5651.44 | 770.597 | 1.39812 | 0.310342 |
| SA 6h | 3802.23 | 532.766 | 1.0723 | 0.0619223 |
| Me-JA 6h | 2608.09 | 783.201 | 0.679925 | 0.156871 |
Source Transcript PGSC0003DMT400084732 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G27950.1 | +1 | 3e-41 | 144 | 70/126 (56%) | glycosylphosphatidylinositol-anchored lipid protein transfer 1 | chr1:9740740-9741991 FORWARD LENGTH=193 |
