Probe CUST_24922_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24922_PI426222305 | JHI_St_60k_v1 | DMT400024088 | TGTCCAGGAATGCATTTTGGTTTAGCTAACGTTGTGTATCCGCTAGCACAGTTGCTTTAT |
All Microarray Probes Designed to Gene DMG400009316
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24922_PI426222305 | JHI_St_60k_v1 | DMT400024088 | TGTCCAGGAATGCATTTTGGTTTAGCTAACGTTGTGTATCCGCTAGCACAGTTGCTTTAT |
Microarray Signals from CUST_24922_PI426222305
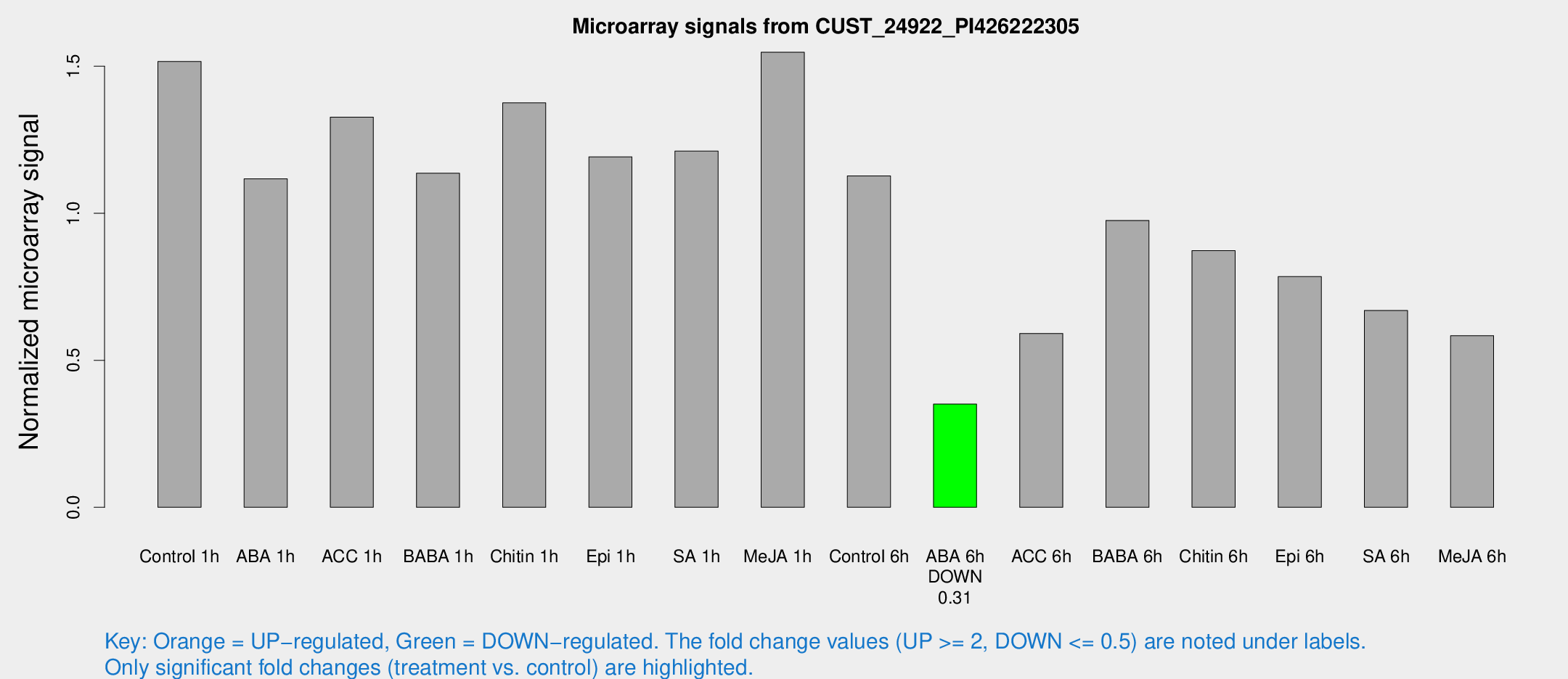
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1375.88 | 129.779 | 1.51619 | 0.0875957 |
| ABA 1h | 909.829 | 143.99 | 1.11696 | 0.116375 |
| ACC 1h | 1233.29 | 98.1168 | 1.32689 | 0.0878288 |
| BABA 1h | 1026.34 | 193.826 | 1.13616 | 0.122534 |
| Chitin 1h | 1136.77 | 162.855 | 1.37571 | 0.167166 |
| Epi 1h | 940.726 | 108.03 | 1.19211 | 0.112654 |
| SA 1h | 1144.6 | 160.423 | 1.2115 | 0.124823 |
| Me-JA 1h | 1137.69 | 65.8118 | 1.54779 | 0.0894495 |
| Control 6h | 1156.74 | 371.006 | 1.12707 | 0.35759 |
| ABA 6h | 336.825 | 19.749 | 0.351105 | 0.0314269 |
| ACC 6h | 648.61 | 163.075 | 0.591307 | 0.0683595 |
| BABA 6h | 985.115 | 57.14 | 0.975576 | 0.0591795 |
| Chitin 6h | 841.733 | 67.5086 | 0.873079 | 0.0521725 |
| Epi 6h | 806.519 | 94.7528 | 0.784557 | 0.0820223 |
| SA 6h | 624.234 | 125.819 | 0.669284 | 0.0739806 |
| Me-JA 6h | 535.209 | 77.0806 | 0.583666 | 0.0473048 |
Source Transcript PGSC0003DMT400024088 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26310.1 | +1 | 9e-98 | 304 | 170/460 (37%) | cytochrome P450, family 71, subfamily B, polypeptide 35 | chr3:9641089-9642779 REVERSE LENGTH=500 |
