Probe CUST_24868_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24868_PI426222305 | JHI_St_60k_v1 | DMT400045229 | TGCAGCAGAGCATGGCTATTTTGTGAGGCCAAATCATGATGCCAAATGGGAAACTTGTGT |
All Microarray Probes Designed to Gene DMG401017546
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24868_PI426222305 | JHI_St_60k_v1 | DMT400045229 | TGCAGCAGAGCATGGCTATTTTGTGAGGCCAAATCATGATGCCAAATGGGAAACTTGTGT |
| CUST_24822_PI426222305 | JHI_St_60k_v1 | DMT400045227 | ATTTGATGGTTTGCTTTTGCCATGAACGTTCATTTCGACCTCCTAGGGTGTCAACAAAGG |
Microarray Signals from CUST_24868_PI426222305
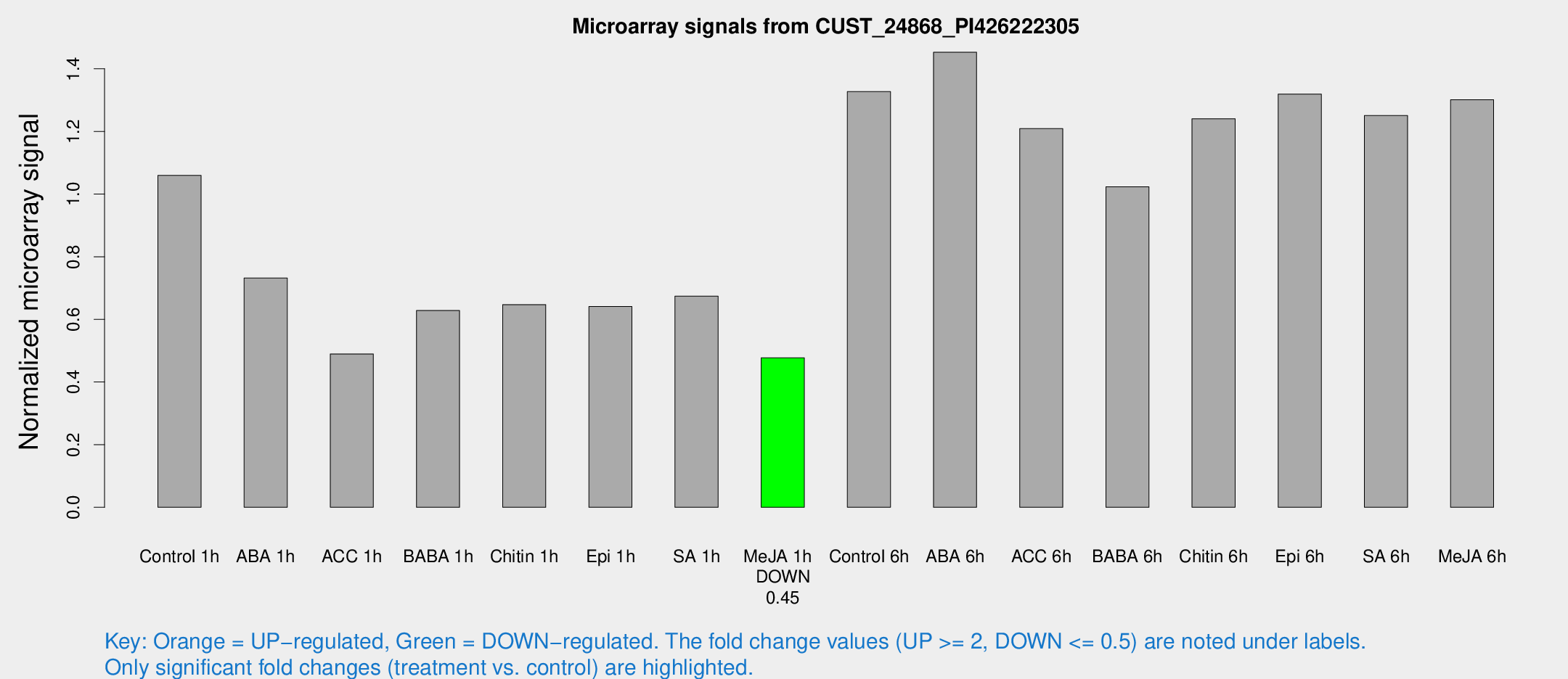
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 845.529 | 57.3427 | 1.0597 | 0.0613193 |
| ABA 1h | 515 | 29.9457 | 0.731858 | 0.0651461 |
| ACC 1h | 488.745 | 171.313 | 0.489686 | 0.237939 |
| BABA 1h | 507.855 | 115.997 | 0.628336 | 0.0951705 |
| Chitin 1h | 473.576 | 76.4155 | 0.647222 | 0.0667874 |
| Epi 1h | 461.812 | 92.8643 | 0.640905 | 0.142213 |
| SA 1h | 565.865 | 95.4245 | 0.674301 | 0.102335 |
| Me-JA 1h | 312.584 | 30.135 | 0.477457 | 0.028045 |
| Control 6h | 1175.99 | 392.265 | 1.32722 | 0.378309 |
| ABA 6h | 1265.47 | 208.797 | 1.45295 | 0.215196 |
| ACC 6h | 1120.05 | 138.098 | 1.20891 | 0.122748 |
| BABA 6h | 938.689 | 173.08 | 1.02329 | 0.17982 |
| Chitin 6h | 1051.66 | 60.9878 | 1.24068 | 0.0717639 |
| Epi 6h | 1200.31 | 144.005 | 1.31933 | 0.0970736 |
| SA 6h | 1039.91 | 240.298 | 1.25074 | 0.179455 |
| Me-JA 6h | 1069.31 | 192.96 | 1.30129 | 0.162839 |
Source Transcript PGSC0003DMT400045229 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G17770.1 | +2 | 0.0 | 1350 | 670/845 (79%) | trehalose phosphatase/synthase 5 | chr4:9877055-9880084 FORWARD LENGTH=862 |
