Probe CUST_24848_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24848_PI426222305 | JHI_St_60k_v1 | DMT400045129 | CGCGAAGTCAATTCTAAGCATAACATTGTTACTGAAGAATGAAGTATGTAGTGATTGATC |
All Microarray Probes Designed to Gene DMG400017505
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24848_PI426222305 | JHI_St_60k_v1 | DMT400045129 | CGCGAAGTCAATTCTAAGCATAACATTGTTACTGAAGAATGAAGTATGTAGTGATTGATC |
| CUST_24840_PI426222305 | JHI_St_60k_v1 | DMT400045130 | CGCGAAGTCAATTCTAAGCATAACATTGTTACTGAAGAATGAAGTATGTAGTGATTGATC |
Microarray Signals from CUST_24848_PI426222305
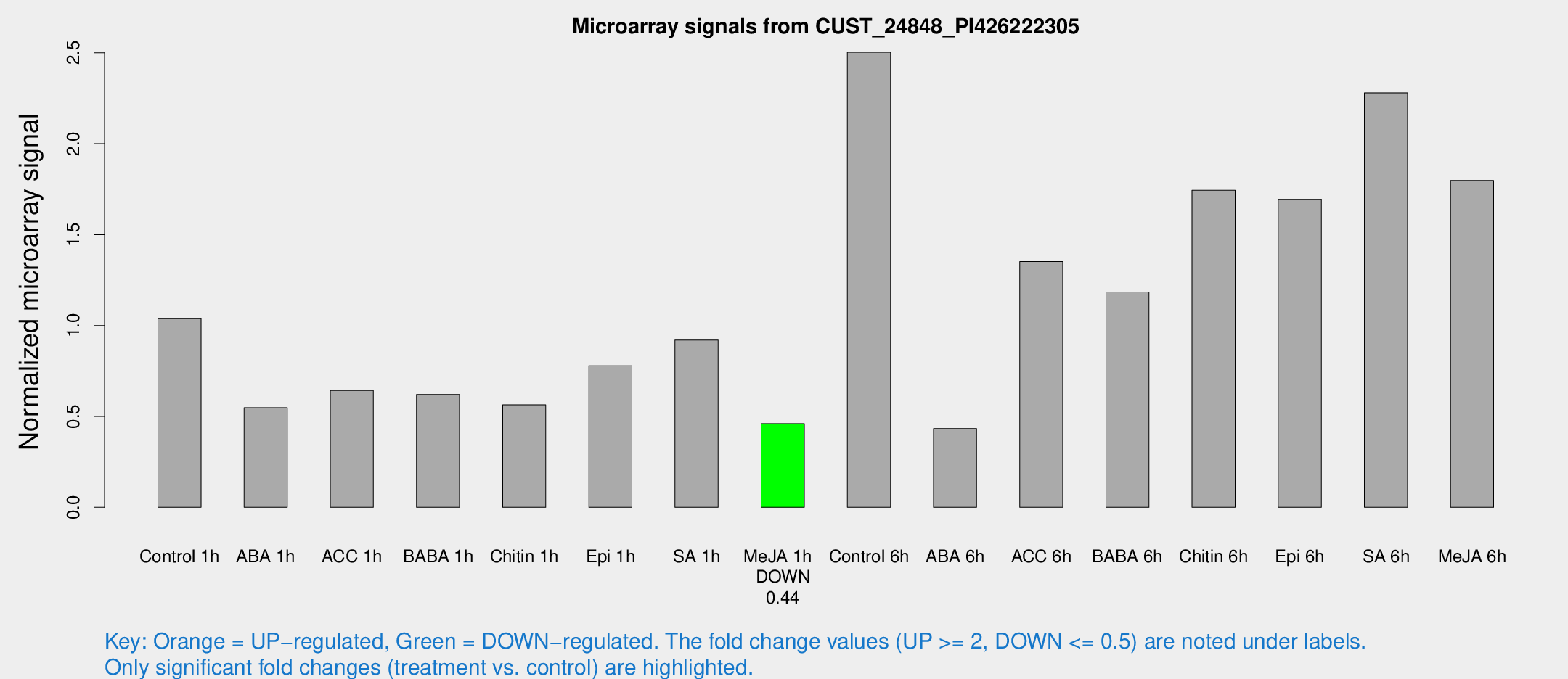
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 109.011 | 8.16275 | 1.03821 | 0.0684849 |
| ABA 1h | 50.6213 | 4.33322 | 0.548036 | 0.0469313 |
| ACC 1h | 78.4274 | 28.6064 | 0.642495 | 0.211818 |
| BABA 1h | 68.4063 | 19.7889 | 0.621213 | 0.138639 |
| Chitin 1h | 56.0615 | 13.4049 | 0.563882 | 0.0924336 |
| Epi 1h | 71.8538 | 10.8554 | 0.778662 | 0.121732 |
| SA 1h | 109.606 | 31.5583 | 0.920047 | 0.295586 |
| Me-JA 1h | 39.8645 | 4.84489 | 0.460217 | 0.0505841 |
| Control 6h | 277.874 | 67.5538 | 2.50303 | 0.470937 |
| ABA 6h | 51.5851 | 14.3684 | 0.433392 | 0.121108 |
| ACC 6h | 167.566 | 31.8331 | 1.35183 | 0.329547 |
| BABA 6h | 157.961 | 52.4468 | 1.1848 | 0.537358 |
| Chitin 6h | 199.022 | 31.5158 | 1.74391 | 0.277323 |
| Epi 6h | 200.395 | 15.7056 | 1.6916 | 0.104038 |
| SA 6h | 245.125 | 45.2388 | 2.27975 | 0.204695 |
| Me-JA 6h | 232.74 | 112.05 | 1.79757 | 0.826046 |
Source Transcript PGSC0003DMT400045129 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G46590.1 | +1 | 3e-95 | 295 | 152/303 (50%) | NAC domain containing protein 96 | chr5:18905679-18906811 FORWARD LENGTH=292 |
