Probe CUST_2480_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2480_PI426222305 | JHI_St_60k_v1 | DMT400027158 | CAAGACACTCATGCCTCGAGAGGGTGTTTTGTCTTATGGAAGGTGTCAAGGAATATTGAT |
All Microarray Probes Designed to Gene DMG400010477
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2476_PI426222305 | JHI_St_60k_v1 | DMT400027157 | AGGGAGAGTGATGTTGTAGTAAAGGGTTCAAAATTCAAGAGGATTTGTGTGTTTTGTGGA |
| CUST_2480_PI426222305 | JHI_St_60k_v1 | DMT400027158 | CAAGACACTCATGCCTCGAGAGGGTGTTTTGTCTTATGGAAGGTGTCAAGGAATATTGAT |
Microarray Signals from CUST_2480_PI426222305
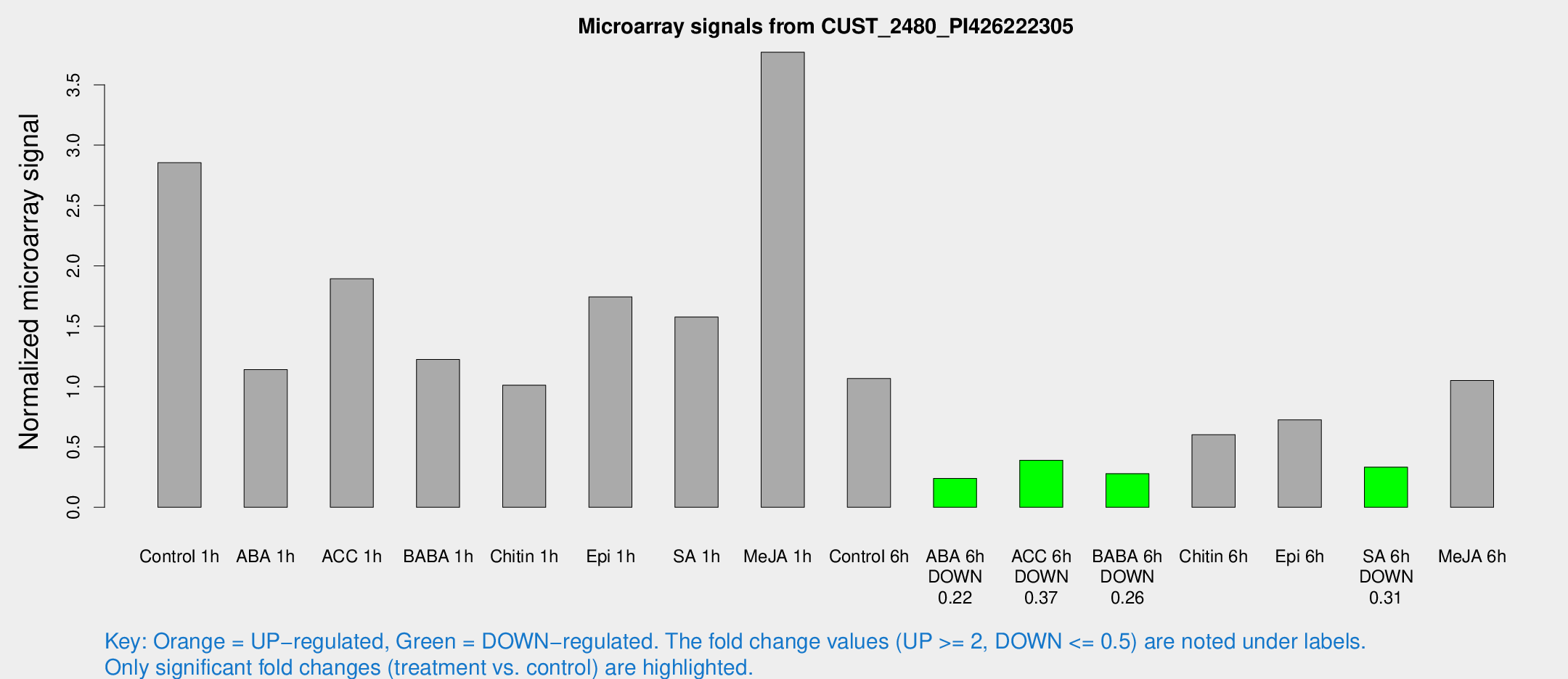
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 74.0628 | 15.8841 | 2.85472 | 0.45597 |
| ABA 1h | 25.9641 | 5.17835 | 1.14105 | 0.213416 |
| ACC 1h | 51.6486 | 12.4164 | 1.89285 | 0.405411 |
| BABA 1h | 32.3845 | 10.7394 | 1.22457 | 0.322415 |
| Chitin 1h | 23.6741 | 5.26815 | 1.01099 | 0.338863 |
| Epi 1h | 37.6167 | 3.99277 | 1.74391 | 0.190649 |
| SA 1h | 42.8844 | 11.7598 | 1.57597 | 0.457357 |
| Me-JA 1h | 79.8611 | 18.3213 | 3.76987 | 0.519469 |
| Control 6h | 26.5877 | 3.86316 | 1.06704 | 0.174434 |
| ABA 6h | 6.30172 | 3.65732 | 0.239587 | 0.138944 |
| ACC 6h | 11.11 | 4.34589 | 0.390339 | 0.152398 |
| BABA 6h | 7.79072 | 3.98223 | 0.278695 | 0.144119 |
| Chitin 6h | 16.9775 | 4.65781 | 0.602125 | 0.179383 |
| Epi 6h | 21.4834 | 4.80992 | 0.723936 | 0.261186 |
| SA 6h | 8.50491 | 3.73545 | 0.332549 | 0.161175 |
| Me-JA 6h | 27.609 | 6.38822 | 1.05101 | 0.219393 |
Source Transcript PGSC0003DMT400027158 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G53450.1 | +3 | 7e-106 | 311 | 155/170 (91%) | Putative lysine decarboxylase family protein | chr3:19812977-19815430 REVERSE LENGTH=215 |
