Probe CUST_24379_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24379_PI426222305 | JHI_St_60k_v1 | DMT400035715 | ACACAGATTCTAACATTAGTTCTAGGTGCTGTTGAGCTTTATATATCTTTTAGACACAGC |
All Microarray Probes Designed to Gene DMG400013736
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24445_PI426222305 | JHI_St_60k_v1 | DMT400035714 | CAGATTCTAACATTAGTTCTAGGTGCTGTTGAGCTTTATATATCTTTTAGACACAGCTTG |
| CUST_24379_PI426222305 | JHI_St_60k_v1 | DMT400035715 | ACACAGATTCTAACATTAGTTCTAGGTGCTGTTGAGCTTTATATATCTTTTAGACACAGC |
Microarray Signals from CUST_24379_PI426222305
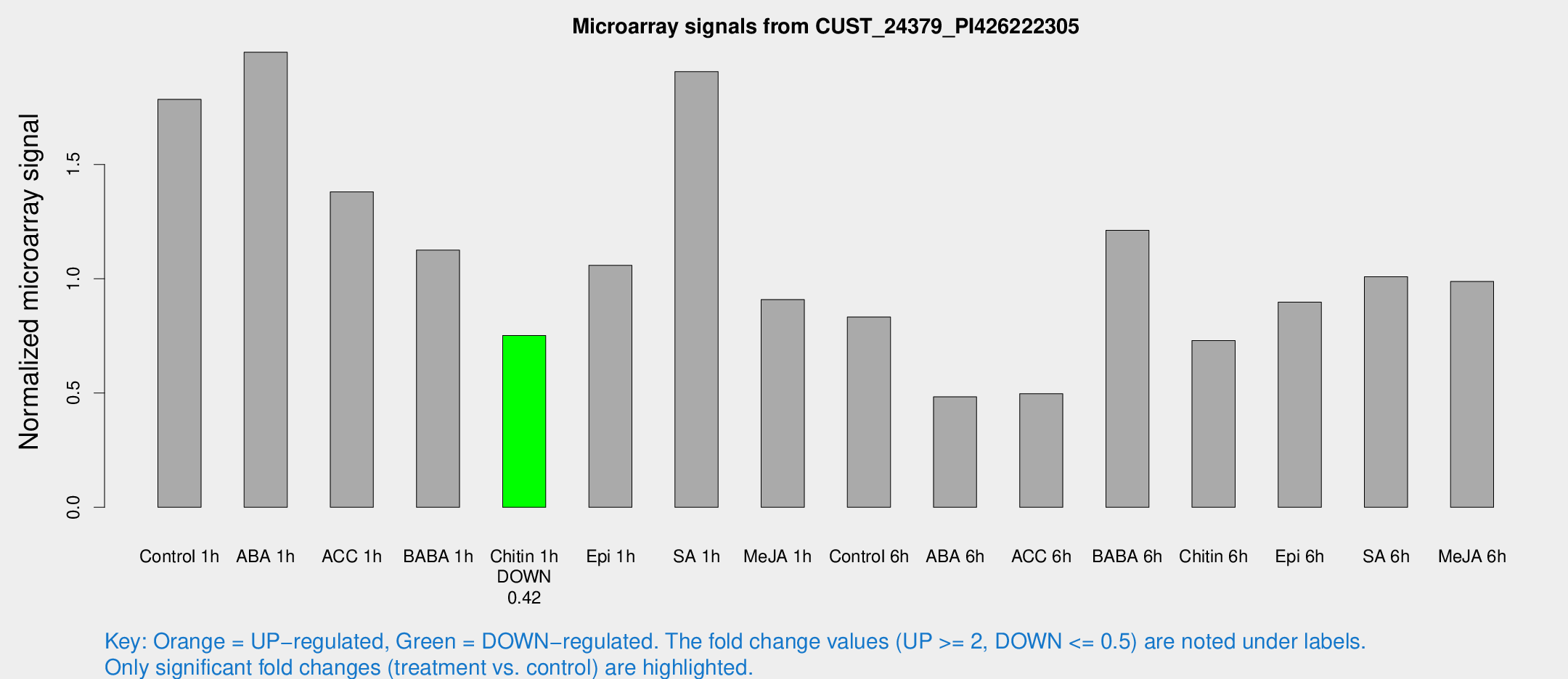
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 72.2055 | 19.258 | 1.78515 | 0.371544 |
| ABA 1h | 67.3422 | 6.52699 | 1.99127 | 0.146628 |
| ACC 1h | 57.6211 | 14.8058 | 1.38004 | 0.279163 |
| BABA 1h | 42.1284 | 6.28811 | 1.12549 | 0.115299 |
| Chitin 1h | 25.832 | 3.4906 | 0.751859 | 0.102366 |
| Epi 1h | 34.873 | 3.71575 | 1.05884 | 0.11295 |
| SA 1h | 76.2052 | 11.7207 | 1.90634 | 0.240088 |
| Me-JA 1h | 29.6376 | 6.5698 | 0.908976 | 0.135866 |
| Control 6h | 33.5671 | 7.4188 | 0.832621 | 0.135421 |
| ABA 6h | 25.4944 | 10.0597 | 0.483343 | 0.337256 |
| ACC 6h | 21.9742 | 4.21933 | 0.497072 | 0.0998429 |
| BABA 6h | 54.2172 | 12.1607 | 1.21168 | 0.25982 |
| Chitin 6h | 29.4622 | 4.09509 | 0.729258 | 0.101834 |
| Epi 6h | 38.7033 | 4.46138 | 0.897842 | 0.146208 |
| SA 6h | 38.5624 | 5.48195 | 1.00915 | 0.164788 |
| Me-JA 6h | 44.4127 | 18.9905 | 0.988109 | 0.356174 |
Source Transcript PGSC0003DMT400035715 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G20830.2 | -3 | 2e-12 | 59 | 45/163 (28%) | transferases;folic acid binding | chr2:8968370-8970479 REVERSE LENGTH=431 |
