Probe CUST_24365_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24365_PI426222305 | JHI_St_60k_v1 | DMT400074120 | ATTGGAGAGACTCACTTTACTCTGTTATGGCTCCTAATCCTGCTAATCCCGAGGAAATCC |
All Microarray Probes Designed to Gene DMG400028809
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24288_PI426222305 | JHI_St_60k_v1 | DMT400074121 | GAAATCCCCGAGACATGCAGGTACTTTTTCACTTATATATACTGACAACGAAAAACAATT |
| CUST_24365_PI426222305 | JHI_St_60k_v1 | DMT400074120 | ATTGGAGAGACTCACTTTACTCTGTTATGGCTCCTAATCCTGCTAATCCCGAGGAAATCC |
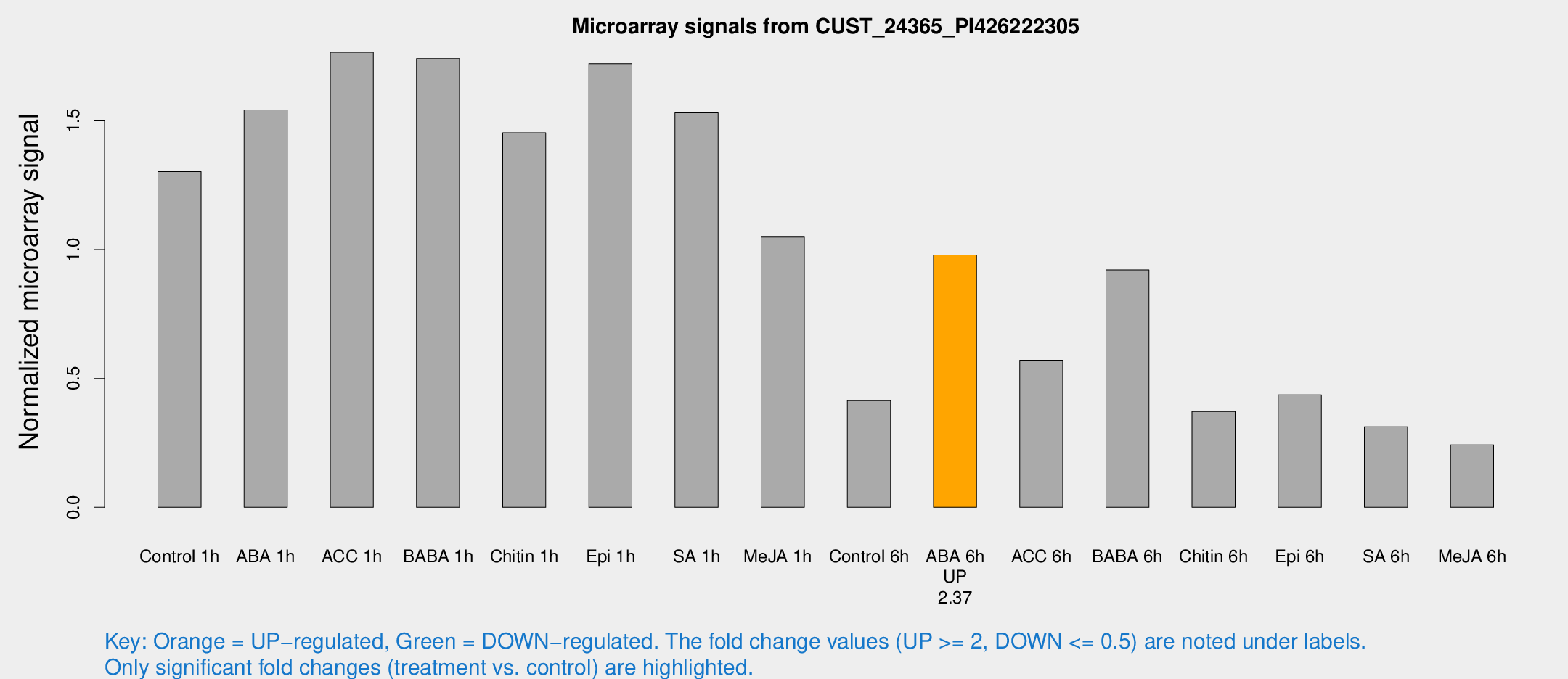
Microarray Signals from CUST_24365_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 913.508 | 99.4147 | 1.30244 | 0.136319 |
| ABA 1h | 948.295 | 57.7195 | 1.54171 | 0.190151 |
| ACC 1h | 1291.87 | 206.073 | 1.76582 | 0.18211 |
| BABA 1h | 1220.14 | 271.913 | 1.74119 | 0.261601 |
| Chitin 1h | 940.063 | 167.456 | 1.45337 | 0.200626 |
| Epi 1h | 1036.12 | 61.5497 | 1.72161 | 0.143679 |
| SA 1h | 1275.29 | 458.084 | 1.53076 | 0.649813 |
| Me-JA 1h | 635.112 | 168.33 | 1.0486 | 0.264264 |
| Control 6h | 313.349 | 80.1626 | 0.413987 | 0.0927286 |
| ABA 6h | 764.863 | 167.135 | 0.97922 | 0.207596 |
| ACC 6h | 459.806 | 54.0282 | 0.571008 | 0.0332519 |
| BABA 6h | 719.129 | 57.1562 | 0.921247 | 0.0761476 |
| Chitin 6h | 275.109 | 16.2681 | 0.371913 | 0.031867 |
| Epi 6h | 363.89 | 93.4917 | 0.43629 | 0.1341 |
| SA 6h | 219.371 | 32.7385 | 0.312965 | 0.0249595 |
| Me-JA 6h | 174.495 | 36.8051 | 0.24256 | 0.0307838 |
Source Transcript PGSC0003DMT400074120 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G06620.1 | +3 | 2e-127 | 379 | 197/356 (55%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:2025618-2027094 FORWARD LENGTH=365 |
