Probe CUST_23981_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23981_PI426222305 | JHI_St_60k_v1 | DMT400032715 | GCTCGTGACCTGTTCCCGTCTTTGTTTTCCCTTTATTCCCCCTTAAAGCTAATTCATAAT |
All Microarray Probes Designed to Gene DMG400012562
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23981_PI426222305 | JHI_St_60k_v1 | DMT400032715 | GCTCGTGACCTGTTCCCGTCTTTGTTTTCCCTTTATTCCCCCTTAAAGCTAATTCATAAT |
| CUST_23859_PI426222305 | JHI_St_60k_v1 | DMT400032713 | AACTTGATCTCTCATGGAGAAATTGCATTTCAAACAGATGGAGAAGCTGCTGATCATTGA |
Microarray Signals from CUST_23981_PI426222305
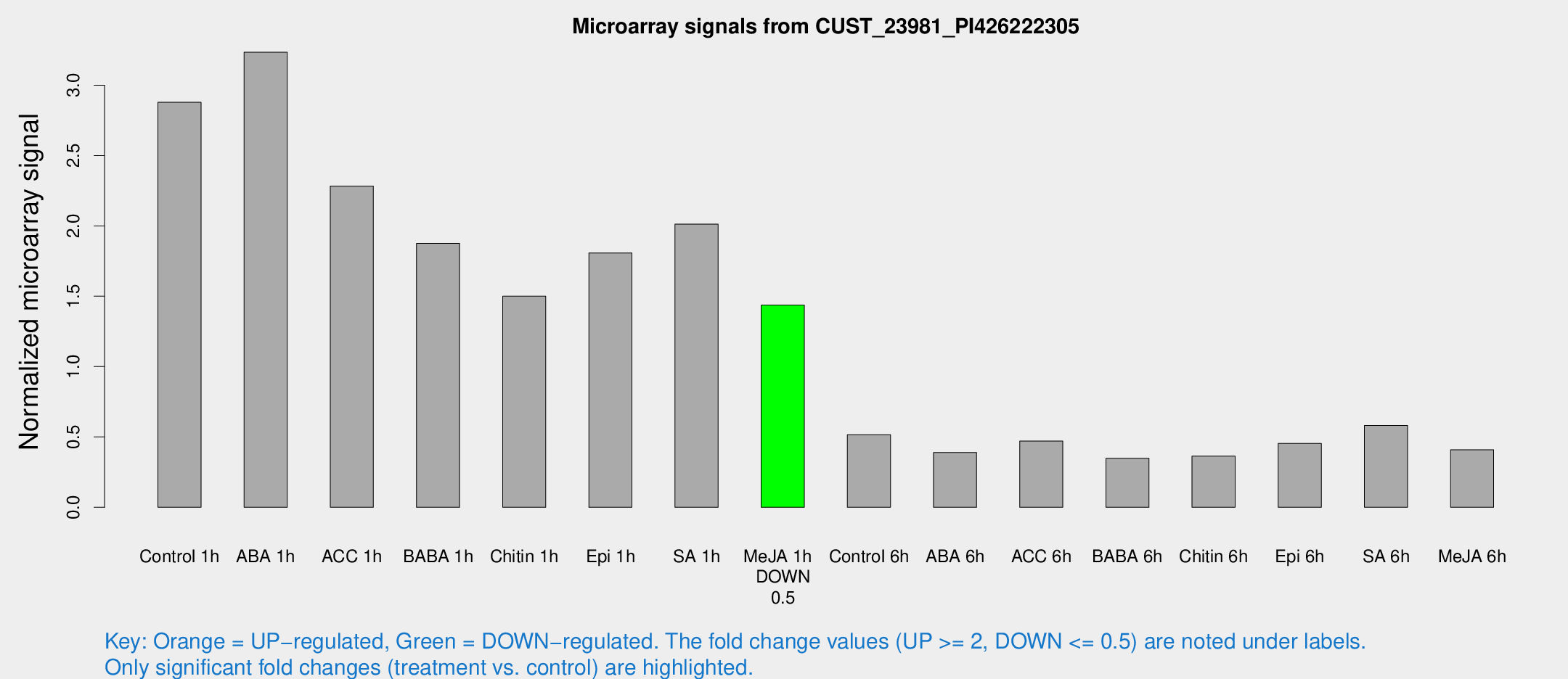
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1429.28 | 158.042 | 2.8786 | 0.172868 |
| ABA 1h | 1407.42 | 81.4915 | 3.23408 | 0.186846 |
| ACC 1h | 1172.1 | 149.681 | 2.28338 | 0.136726 |
| BABA 1h | 918.431 | 165.574 | 1.87572 | 0.187298 |
| Chitin 1h | 667.924 | 64.1014 | 1.5 | 0.162871 |
| Epi 1h | 779.79 | 94.4642 | 1.80792 | 0.218914 |
| SA 1h | 1017.65 | 58.9991 | 2.01243 | 0.116351 |
| Me-JA 1h | 578.017 | 33.6253 | 1.43809 | 0.083398 |
| Control 6h | 262.194 | 48.4135 | 0.516193 | 0.0756655 |
| ABA 6h | 203.695 | 12.2316 | 0.389411 | 0.0262445 |
| ACC 6h | 266.21 | 19.4523 | 0.470303 | 0.0393309 |
| BABA 6h | 192.839 | 14.7345 | 0.34886 | 0.0367472 |
| Chitin 6h | 190.789 | 11.5789 | 0.364311 | 0.0220824 |
| Epi 6h | 252.367 | 17.6624 | 0.453819 | 0.035418 |
| SA 6h | 294.103 | 62.5398 | 0.581201 | 0.0946342 |
| Me-JA 6h | 205.42 | 34.8387 | 0.408427 | 0.0352756 |
Source Transcript PGSC0003DMT400032715 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G64170.2 | +1 | 7e-71 | 249 | 211/669 (32%) | dentin sialophosphoprotein-related | chr5:25672904-25675934 REVERSE LENGTH=616 |
