Probe CUST_23956_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23956_PI426222305 | JHI_St_60k_v1 | DMT400076573 | GTTGTTGTAATGCTTCTAATTAGTTAGTCGCCCTGTAATTTTTCCAAGTTATGGCTTTGT |
All Microarray Probes Designed to Gene DMG400029773
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23956_PI426222305 | JHI_St_60k_v1 | DMT400076573 | GTTGTTGTAATGCTTCTAATTAGTTAGTCGCCCTGTAATTTTTCCAAGTTATGGCTTTGT |
| CUST_23876_PI426222305 | JHI_St_60k_v1 | DMT400076572 | GAAATGGAAAGGCTATAGTTGTTCCTTGTATAGATGTGAAGGATCTCAGTGTTTTCTAAT |
Microarray Signals from CUST_23956_PI426222305
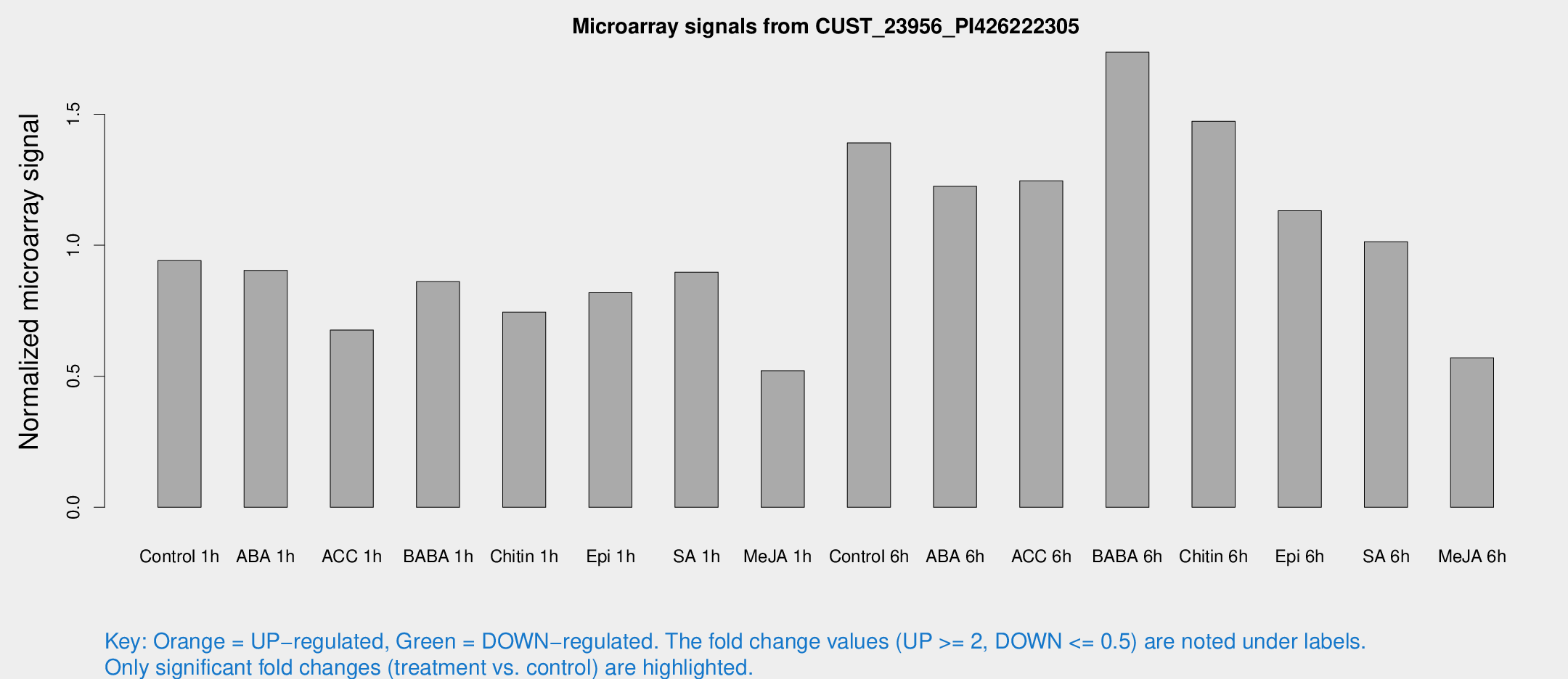
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 208.796 | 20.6069 | 0.941389 | 0.0778271 |
| ABA 1h | 189.971 | 52.8926 | 0.904308 | 0.195933 |
| ACC 1h | 157.393 | 28.6345 | 0.67652 | 0.120353 |
| BABA 1h | 197.403 | 49.387 | 0.861357 | 0.171023 |
| Chitin 1h | 151.515 | 24.1116 | 0.745022 | 0.144947 |
| Epi 1h | 162.024 | 32.03 | 0.818933 | 0.147861 |
| SA 1h | 216.279 | 58.3235 | 0.897409 | 0.185994 |
| Me-JA 1h | 100.387 | 27.2014 | 0.52118 | 0.119997 |
| Control 6h | 319.851 | 68.2529 | 1.39034 | 0.225839 |
| ABA 6h | 298.5 | 58.7476 | 1.22529 | 0.194801 |
| ACC 6h | 336.234 | 80.5618 | 1.24566 | 0.235813 |
| BABA 6h | 430.029 | 35.1422 | 1.73647 | 0.101325 |
| Chitin 6h | 366.562 | 93.7881 | 1.47307 | 0.344006 |
| Epi 6h | 283.853 | 30.9825 | 1.13195 | 0.0670724 |
| SA 6h | 241.643 | 65.788 | 1.01362 | 0.216837 |
| Me-JA 6h | 142.506 | 43.3322 | 0.570749 | 0.193529 |
Source Transcript PGSC0003DMT400076573 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G24500.1 | +1 | 8e-53 | 174 | 98/143 (69%) | multiprotein bridging factor 1C | chr3:8918755-8919201 FORWARD LENGTH=148 |
