Probe CUST_23818_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23818_PI426222305 | JHI_St_60k_v1 | DMT400021173 | GCCTATTGAAGCTGAGACAATCGAAGAGGCATTACCGGAGGTTTCAAAGAGAGAACACAA |
All Microarray Probes Designed to Gene DMG400008201
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23818_PI426222305 | JHI_St_60k_v1 | DMT400021173 | GCCTATTGAAGCTGAGACAATCGAAGAGGCATTACCGGAGGTTTCAAAGAGAGAACACAA |
Microarray Signals from CUST_23818_PI426222305
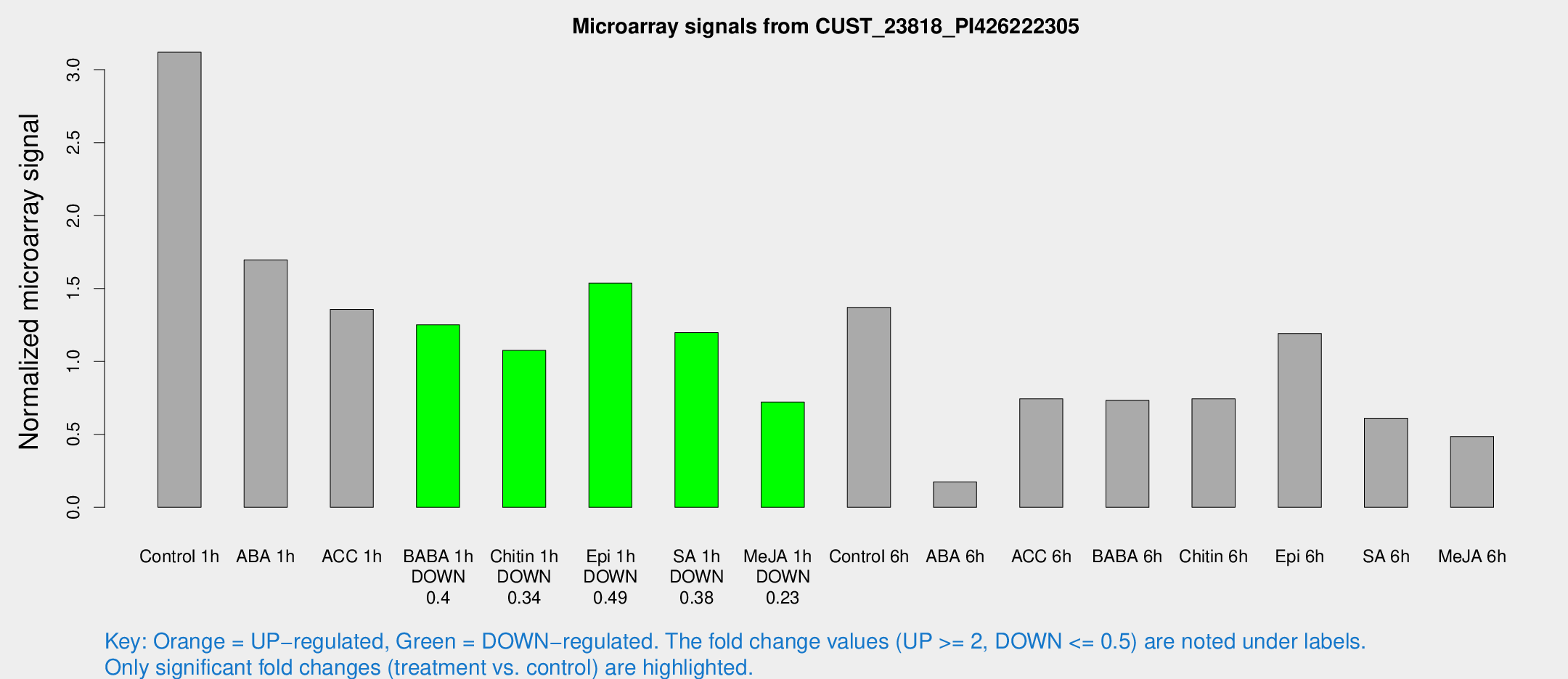
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 822.125 | 155.445 | 3.11982 | 0.402457 |
| ABA 1h | 381.338 | 22.318 | 1.69609 | 0.0994067 |
| ACC 1h | 407.339 | 125.623 | 1.35735 | 0.485977 |
| BABA 1h | 328.597 | 79.5475 | 1.25095 | 0.225858 |
| Chitin 1h | 249.311 | 28.9089 | 1.07572 | 0.0740345 |
| Epi 1h | 345.445 | 48.2576 | 1.53743 | 0.216414 |
| SA 1h | 320.041 | 46.9522 | 1.19808 | 0.143571 |
| Me-JA 1h | 151.286 | 16.2778 | 0.721736 | 0.0454306 |
| Control 6h | 381.09 | 118.089 | 1.37035 | 0.359538 |
| ABA 6h | 47.9862 | 6.4793 | 0.174129 | 0.0356806 |
| ACC 6h | 227.867 | 52.9852 | 0.743726 | 0.0627341 |
| BABA 6h | 218.726 | 47.6529 | 0.732886 | 0.148775 |
| Chitin 6h | 203.204 | 20.8797 | 0.743592 | 0.0546625 |
| Epi 6h | 345.876 | 40.2841 | 1.19195 | 0.188416 |
| SA 6h | 178.81 | 63.4579 | 0.611449 | 0.19547 |
| Me-JA 6h | 133.856 | 40.4021 | 0.485737 | 0.105077 |
Source Transcript PGSC0003DMT400021173 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27570.1 | +1 | 3e-35 | 130 | 60/129 (47%) | UDP-Glycosyltransferase superfamily protein | chr4:13763657-13765018 REVERSE LENGTH=453 |
