Probe CUST_23709_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23709_PI426222305 | JHI_St_60k_v1 | DMT400002375 | AGAGAGAAGTCATATTTTCTGGACTCTCCCATGTGAGTTCCTAGAAAATTCTTGTTAAAG |
All Microarray Probes Designed to Gene DMG400000910
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23709_PI426222305 | JHI_St_60k_v1 | DMT400002375 | AGAGAGAAGTCATATTTTCTGGACTCTCCCATGTGAGTTCCTAGAAAATTCTTGTTAAAG |
Microarray Signals from CUST_23709_PI426222305
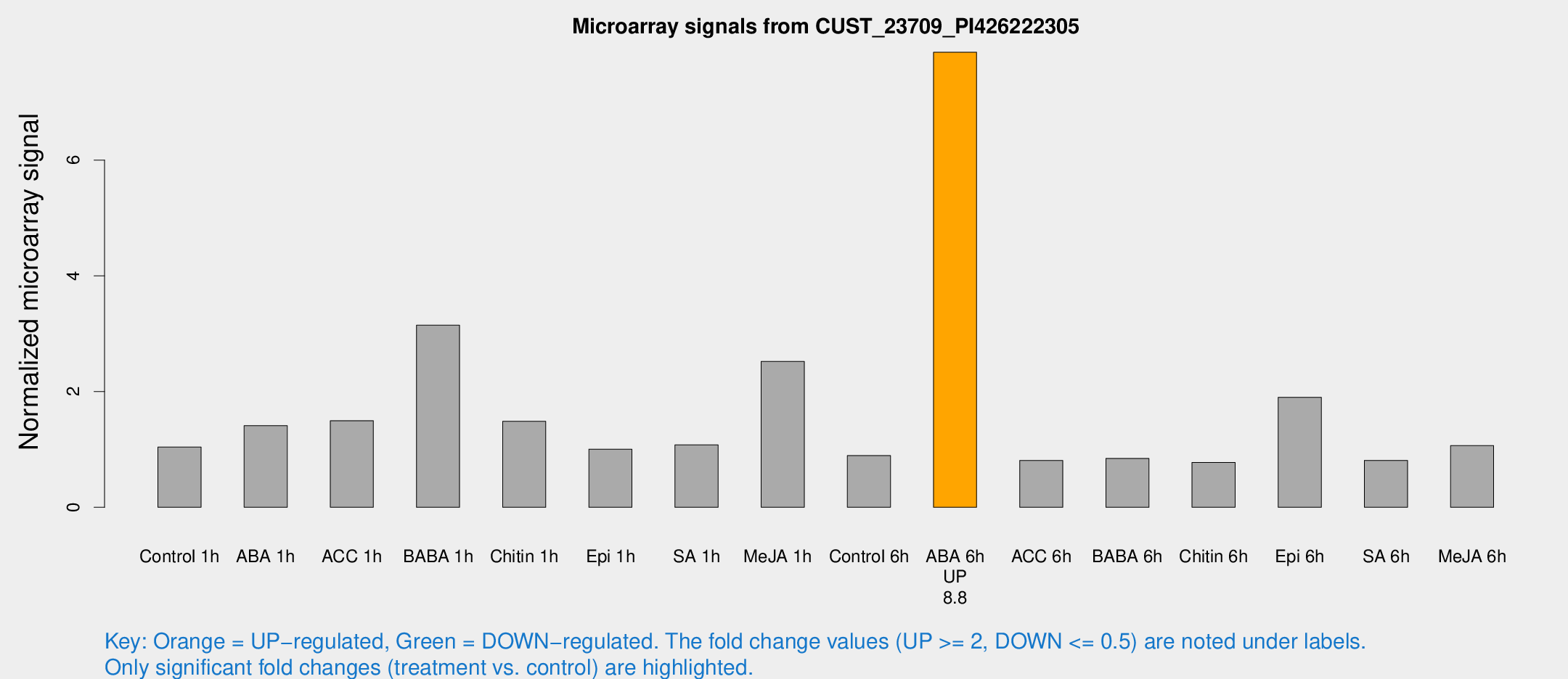
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.81663 | 3.02662 | 1.03997 | 0.472206 |
| ABA 1h | 8.95858 | 2.75794 | 1.41014 | 0.515658 |
| ACC 1h | 12.4182 | 5.11117 | 1.49586 | 0.730277 |
| BABA 1h | 22.1267 | 6.36564 | 3.14573 | 1.32022 |
| Chitin 1h | 10.3703 | 4.49816 | 1.4857 | 0.629833 |
| Epi 1h | 5.88636 | 2.82952 | 1.00464 | 0.486975 |
| SA 1h | 8.8443 | 3.97853 | 1.07927 | 0.488208 |
| Me-JA 1h | 16.5495 | 5.51747 | 2.52051 | 1.71901 |
| Control 6h | 6.11728 | 2.91823 | 0.893608 | 0.435884 |
| ABA 6h | 57.1755 | 6.39798 | 7.8632 | 0.62326 |
| ACC 6h | 6.34597 | 3.54164 | 0.809382 | 0.439001 |
| BABA 6h | 6.58054 | 3.29562 | 0.844736 | 0.440972 |
| Chitin 6h | 5.5821 | 3.23843 | 0.776145 | 0.450191 |
| Epi 6h | 16.7225 | 5.66334 | 1.90125 | 1.00375 |
| SA 6h | 5.42386 | 3.10141 | 0.810914 | 0.463624 |
| Me-JA 6h | 8.01554 | 3.02352 | 1.06547 | 0.490877 |
Source Transcript PGSC0003DMT400002375 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G44830.1 | +2 | 4e-31 | 119 | 100/197 (51%) | Integrase-type DNA-binding superfamily protein | chr1:16933792-16934427 FORWARD LENGTH=211 |
