Probe CUST_23667_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23667_PI426222305 | JHI_St_60k_v1 | DMT400002293 | TTGGGTTTTCTCCATCTTTTCCCATGAATCCAGCTGATTTCCTCCTTGATCTCGCTAATG |
All Microarray Probes Designed to Gene DMG400000879
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23717_PI426222305 | JHI_St_60k_v1 | DMT400002294 | GGCCAGAATTCATTGTTGTAACTCATATTGAGTACAATTAATGGTACTCAAAAACAGGAG |
| CUST_23667_PI426222305 | JHI_St_60k_v1 | DMT400002293 | TTGGGTTTTCTCCATCTTTTCCCATGAATCCAGCTGATTTCCTCCTTGATCTCGCTAATG |
Microarray Signals from CUST_23667_PI426222305
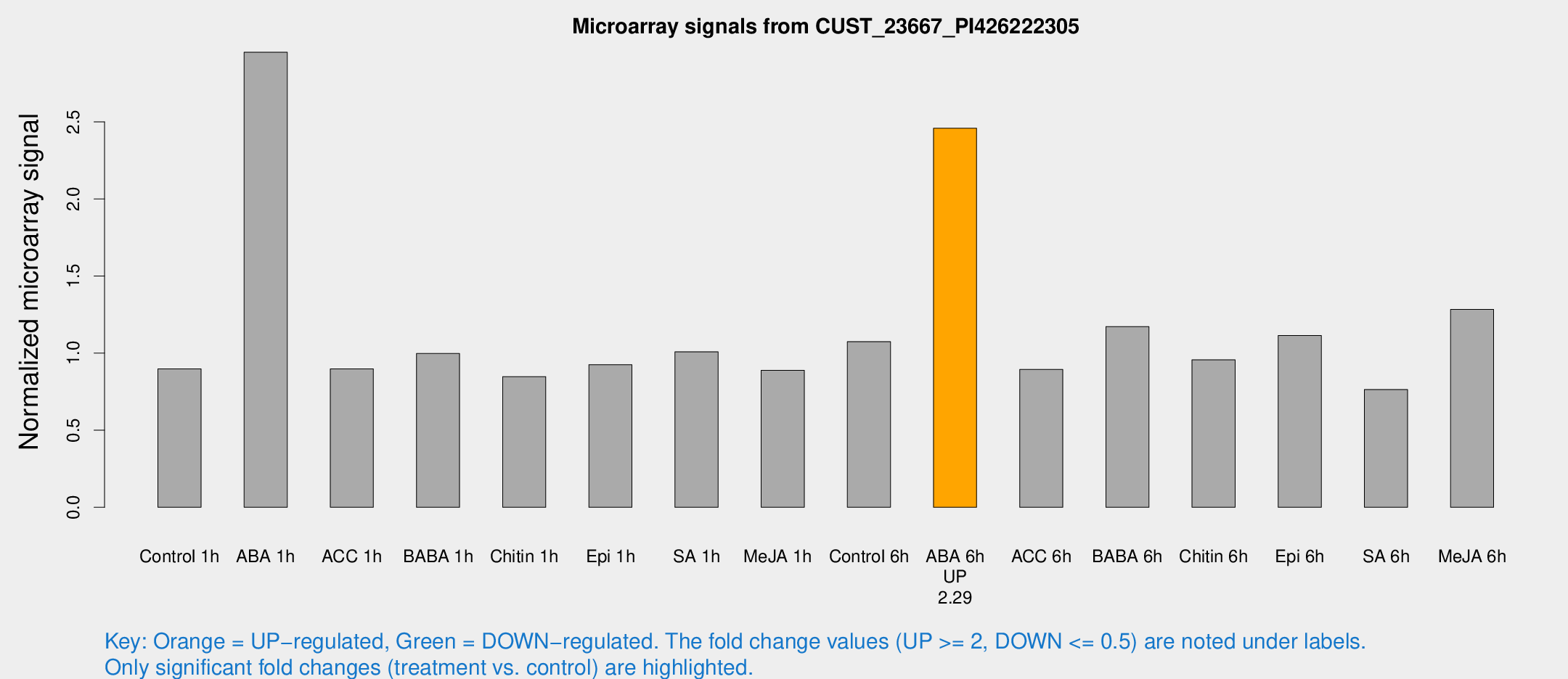
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 187.988 | 22.4399 | 0.898189 | 0.0953449 |
| ABA 1h | 547.245 | 71.0291 | 2.95183 | 0.451566 |
| ACC 1h | 203.986 | 50.1805 | 0.897463 | 0.189782 |
| BABA 1h | 201.662 | 25.7117 | 0.997626 | 0.0616492 |
| Chitin 1h | 157.579 | 9.69122 | 0.847653 | 0.0520118 |
| Epi 1h | 170.872 | 29.2143 | 0.925251 | 0.168313 |
| SA 1h | 214.002 | 12.7858 | 1.00883 | 0.0866239 |
| Me-JA 1h | 151.635 | 17.4493 | 0.888972 | 0.0716236 |
| Control 6h | 242.542 | 62.4271 | 1.07466 | 0.244292 |
| ABA 6h | 561.3 | 102.956 | 2.45872 | 0.41501 |
| ACC 6h | 216.108 | 32.0652 | 0.894502 | 0.0544614 |
| BABA 6h | 272.064 | 18.8656 | 1.1723 | 0.0697279 |
| Chitin 6h | 210.629 | 12.8147 | 0.956248 | 0.0810869 |
| Epi 6h | 272.337 | 63.2482 | 1.11453 | 0.265111 |
| SA 6h | 156.974 | 12.3696 | 0.764212 | 0.047779 |
| Me-JA 6h | 269.303 | 36.9771 | 1.28344 | 0.107956 |
Source Transcript PGSC0003DMT400002293 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G71960.1 | +2 | 0.0 | 761 | 398/621 (64%) | ATP-binding casette family G25 | chr1:27082587-27088163 REVERSE LENGTH=662 |
