Probe CUST_23655_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23655_PI426222305 | JHI_St_60k_v1 | DMT400074655 | GTGTCCCATGATGATAGGGGTTTGGAAAATAAAAACGTATTGTTGAGACTTGAACTTCTA |
All Microarray Probes Designed to Gene DMG400029026
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23655_PI426222305 | JHI_St_60k_v1 | DMT400074655 | GTGTCCCATGATGATAGGGGTTTGGAAAATAAAAACGTATTGTTGAGACTTGAACTTCTA |
Microarray Signals from CUST_23655_PI426222305
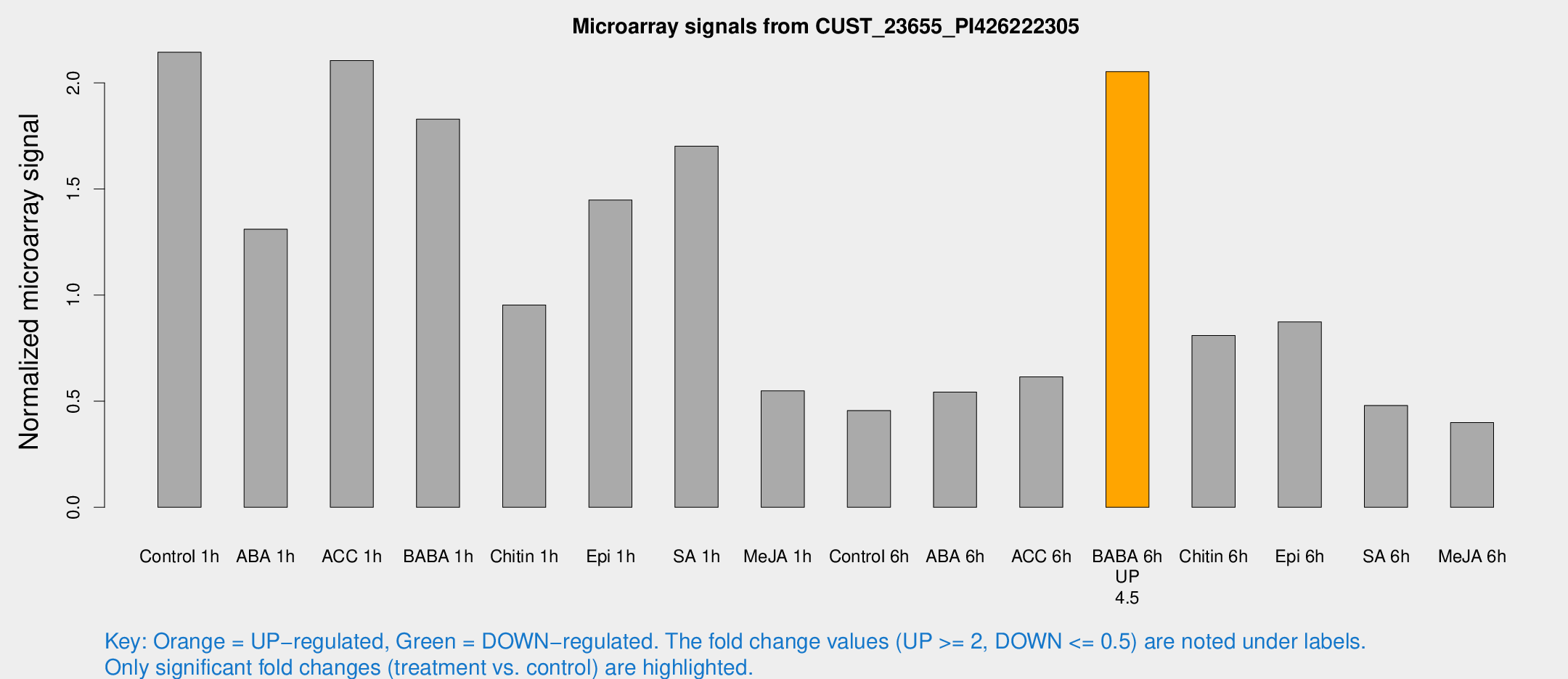
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 54.176 | 13.2003 | 2.14451 | 0.38367 |
| ABA 1h | 27.6612 | 3.14397 | 1.31066 | 0.149478 |
| ACC 1h | 53.4411 | 9.44943 | 2.10556 | 0.258491 |
| BABA 1h | 43.8615 | 8.35089 | 1.82925 | 0.203804 |
| Chitin 1h | 21.4233 | 4.82795 | 0.95341 | 0.164297 |
| Epi 1h | 30.0623 | 3.30884 | 1.44829 | 0.160137 |
| SA 1h | 44.6709 | 11.1423 | 1.70145 | 0.465357 |
| Me-JA 1h | 11.8809 | 3.41554 | 0.548736 | 0.204412 |
| Control 6h | 11.5282 | 2.98663 | 0.455996 | 0.136836 |
| ABA 6h | 13.794 | 3.13992 | 0.543128 | 0.123744 |
| ACC 6h | 17.2269 | 3.6793 | 0.614713 | 0.154343 |
| BABA 6h | 55.0766 | 4.58199 | 2.05275 | 0.171251 |
| Chitin 6h | 20.9289 | 3.46093 | 0.809281 | 0.139708 |
| Epi 6h | 23.808 | 3.64507 | 0.873471 | 0.138692 |
| SA 6h | 11.9498 | 3.1773 | 0.479911 | 0.159598 |
| Me-JA 6h | 12.0467 | 6.22605 | 0.399214 | 0.185247 |
Source Transcript PGSC0003DMT400074655 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G09070.1 | +2 | 6e-18 | 81 | 45/140 (32%) | soybean gene regulated by cold-2 | chr1:2927767-2928741 FORWARD LENGTH=324 |
