Probe CUST_23451_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23451_PI426222305 | JHI_St_60k_v1 | DMT400064924 | GCTTGGTAGCAAGAAGGTGTAGATGGCTGATATACAAAACAAACTTCTAAATGAAAATGT |
All Microarray Probes Designed to Gene DMG400025214
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23451_PI426222305 | JHI_St_60k_v1 | DMT400064924 | GCTTGGTAGCAAGAAGGTGTAGATGGCTGATATACAAAACAAACTTCTAAATGAAAATGT |
Microarray Signals from CUST_23451_PI426222305
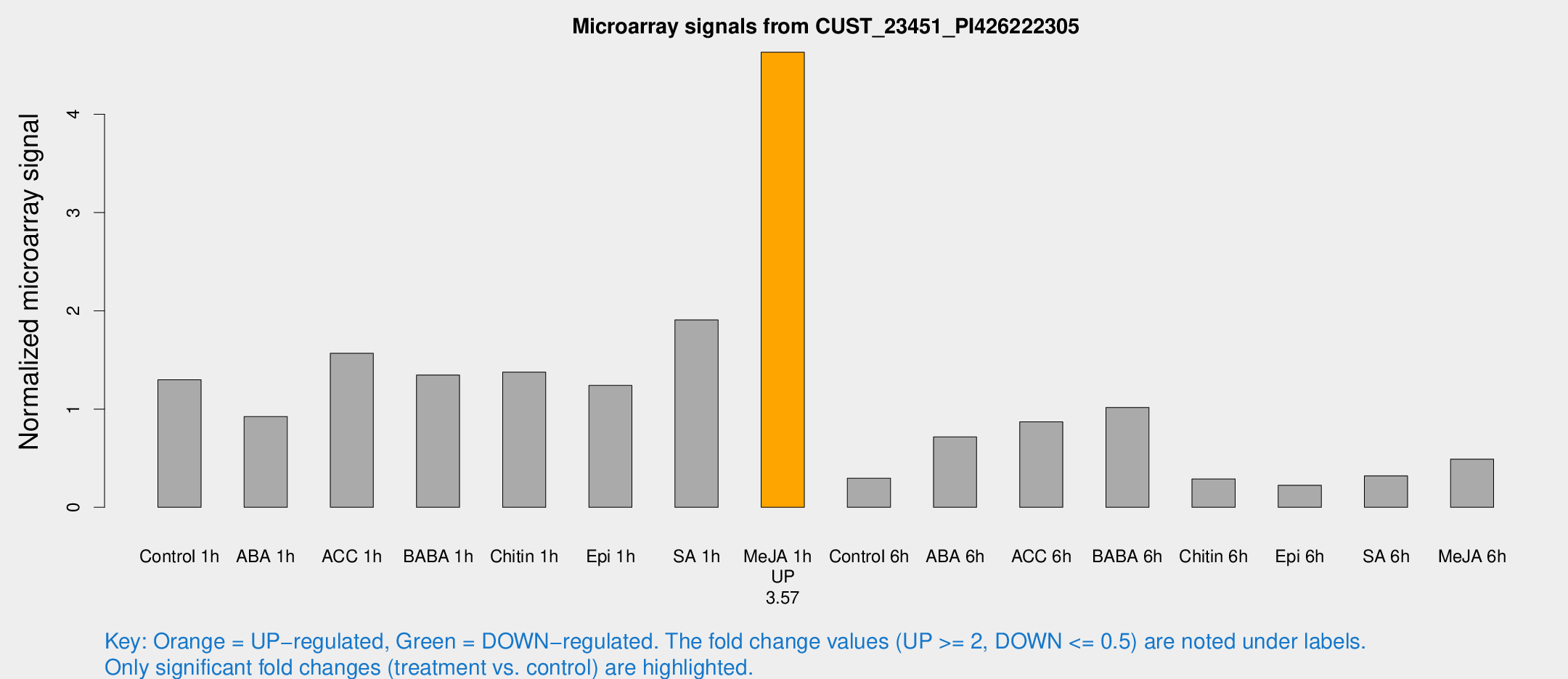
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 532.874 | 77.2007 | 1.29885 | 0.152893 |
| ABA 1h | 331.895 | 36.3798 | 0.9228 | 0.0716543 |
| ACC 1h | 680.473 | 152.196 | 1.56745 | 0.273191 |
| BABA 1h | 552.134 | 119.825 | 1.34626 | 0.198317 |
| Chitin 1h | 498.966 | 30.573 | 1.37536 | 0.0798602 |
| Epi 1h | 431.735 | 25.1365 | 1.2402 | 0.0721471 |
| SA 1h | 849.788 | 222.196 | 1.90665 | 0.498961 |
| Me-JA 1h | 1556.83 | 229.974 | 4.6314 | 0.382915 |
| Control 6h | 131.179 | 36.9273 | 0.295134 | 0.0722206 |
| ABA 6h | 335.466 | 90.8543 | 0.71616 | 0.203105 |
| ACC 6h | 409.642 | 58.2914 | 0.869756 | 0.181514 |
| BABA 6h | 516.62 | 185.304 | 1.01643 | 0.356595 |
| Chitin 6h | 124.044 | 7.97415 | 0.289358 | 0.0185808 |
| Epi 6h | 104.133 | 15.6276 | 0.22398 | 0.0231671 |
| SA 6h | 130.943 | 22.0251 | 0.320138 | 0.0282071 |
| Me-JA 6h | 198.513 | 20.301 | 0.490486 | 0.0673299 |
Source Transcript PGSC0003DMT400064924 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G31820.1 | +2 | 0.0 | 652 | 387/568 (68%) | Ankyrin repeat family protein | chr2:13530350-13532562 FORWARD LENGTH=662 |
