Probe CUST_23440_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23440_PI426222305 | JHI_St_60k_v1 | DMT400064953 | CGATAGAGGGGTATAAAGTCACCCACCCATTTTTTGTAGAATCATATCTTAAATTTCTGT |
All Microarray Probes Designed to Gene DMG400025226
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23440_PI426222305 | JHI_St_60k_v1 | DMT400064953 | CGATAGAGGGGTATAAAGTCACCCACCCATTTTTTGTAGAATCATATCTTAAATTTCTGT |
Microarray Signals from CUST_23440_PI426222305
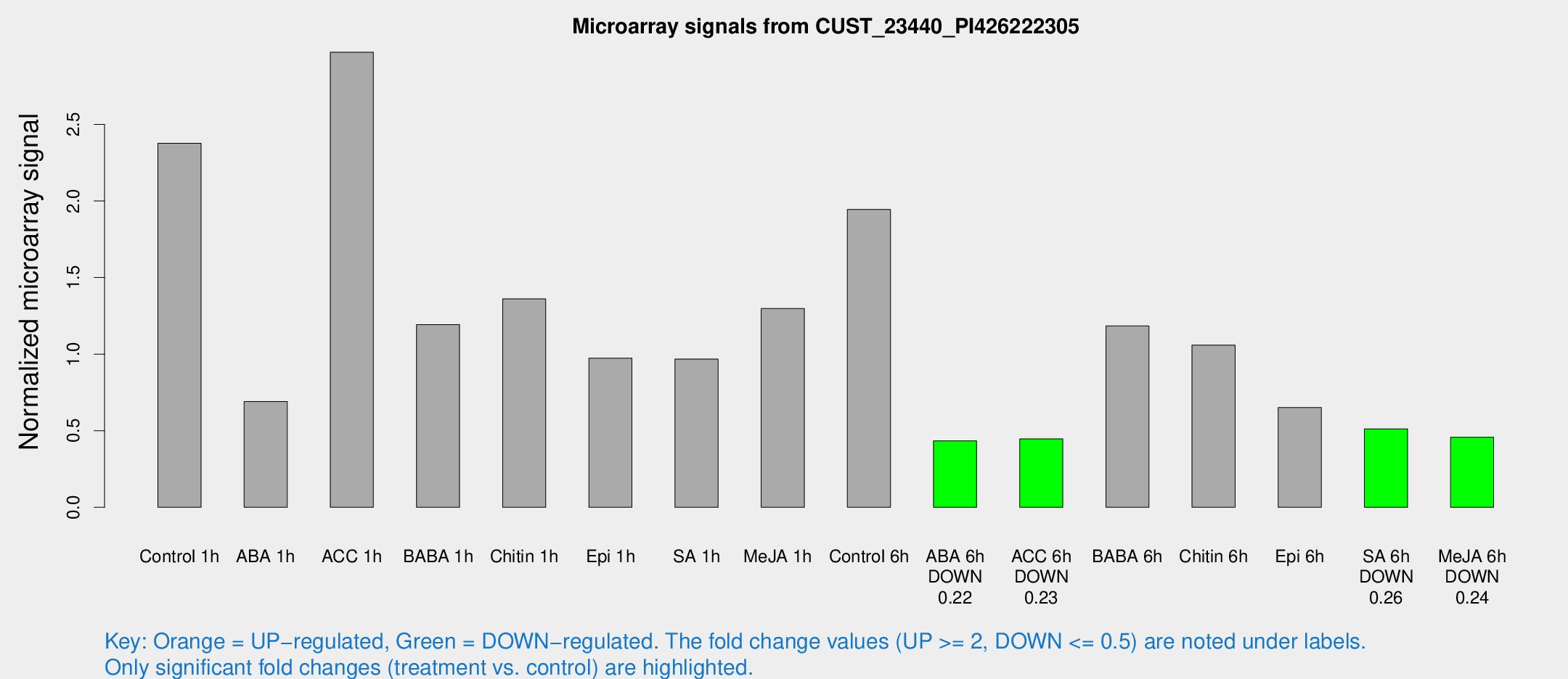
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 33.8741 | 5.84547 | 2.37693 | 0.471964 |
| ABA 1h | 9.75705 | 3.95212 | 0.690034 | 0.342728 |
| ACC 1h | 44.6348 | 11.3145 | 2.97069 | 1.01824 |
| BABA 1h | 15.915 | 3.66413 | 1.19252 | 0.276075 |
| Chitin 1h | 20.3529 | 9.05929 | 1.36058 | 0.724971 |
| Epi 1h | 12.2254 | 3.38396 | 0.973957 | 0.294573 |
| SA 1h | 16.4472 | 5.88308 | 0.967844 | 0.468492 |
| Me-JA 1h | 14.7069 | 3.59592 | 1.29772 | 0.32786 |
| Control 6h | 31.4189 | 11.3995 | 1.94446 | 0.714411 |
| ABA 6h | 6.35512 | 3.68253 | 0.433841 | 0.25122 |
| ACC 6h | 7.2029 | 4.26838 | 0.446185 | 0.258459 |
| BABA 6h | 21.3966 | 7.21664 | 1.18373 | 0.639008 |
| Chitin 6h | 15.593 | 4.03066 | 1.0592 | 0.276786 |
| Epi 6h | 11.6454 | 4.50593 | 0.651047 | 0.302735 |
| SA 6h | 6.99143 | 3.77336 | 0.511403 | 0.278352 |
| Me-JA 6h | 6.27967 | 3.65265 | 0.457461 | 0.265742 |
Source Transcript PGSC0003DMT400064953 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G43900.1 | +2 | 0.0 | 1458 | 732/1131 (65%) | Endonuclease/exonuclease/phosphatase family protein | chr2:18178801-18183823 REVERSE LENGTH=1316 |
