Probe CUST_23289_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23289_PI426222305 | JHI_St_60k_v1 | DMT400002568 | GGTCCCAAAGCGGATACTAGATATATATACTGGATGGGAAACATTAAAAAATGTGTCATT |
All Microarray Probes Designed to Gene DMG400000984
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23289_PI426222305 | JHI_St_60k_v1 | DMT400002568 | GGTCCCAAAGCGGATACTAGATATATATACTGGATGGGAAACATTAAAAAATGTGTCATT |
| CUST_23353_PI426222305 | JHI_St_60k_v1 | DMT400002567 | TTAAGGTTTGTTTTGGTGTGTTCTGCTTTATCCATACATTTCTTCAAGAGAGTGTTAGAG |
Microarray Signals from CUST_23289_PI426222305
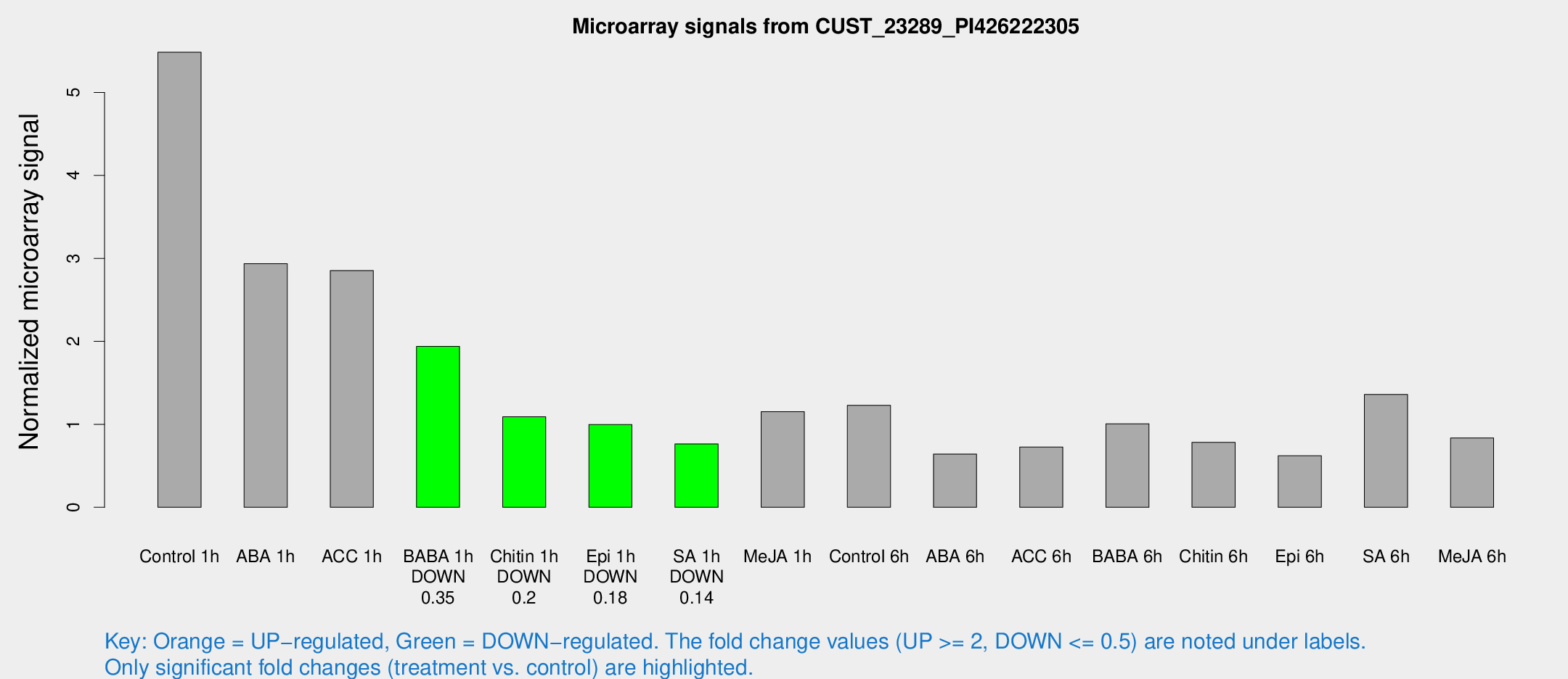
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 52.0623 | 7.96905 | 5.48424 | 0.575322 |
| ABA 1h | 29.9488 | 12.378 | 2.93598 | 2.06706 |
| ACC 1h | 33.0124 | 11.5058 | 2.85256 | 1.35782 |
| BABA 1h | 17.3941 | 3.57612 | 1.93866 | 0.406851 |
| Chitin 1h | 9.3481 | 3.26044 | 1.09038 | 0.398386 |
| Epi 1h | 9.31116 | 3.81055 | 0.997808 | 0.464918 |
| SA 1h | 7.38859 | 3.2098 | 0.763955 | 0.346931 |
| Me-JA 1h | 9.93523 | 3.80038 | 1.15268 | 0.566056 |
| Control 6h | 18.194 | 12.3119 | 1.22968 | 1.08354 |
| ABA 6h | 6.35395 | 3.47368 | 0.642131 | 0.353575 |
| ACC 6h | 7.86052 | 4.08659 | 0.725317 | 0.374595 |
| BABA 6h | 10.86 | 3.73944 | 1.00676 | 0.392186 |
| Chitin 6h | 7.94521 | 3.72507 | 0.782565 | 0.388086 |
| Epi 6h | 6.5317 | 3.81705 | 0.62119 | 0.359987 |
| SA 6h | 14.239 | 4.73276 | 1.36019 | 0.658175 |
| Me-JA 6h | 8.55996 | 3.32282 | 0.835806 | 0.396992 |
Source Transcript PGSC0003DMT400002568 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G16010.1 | +2 | 4e-64 | 208 | 97/157 (62%) | 3-oxo-5-alpha-steroid 4-dehydrogenase family protein | chr5:5227982-5229012 FORWARD LENGTH=268 |
