Probe CUST_23278_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23278_PI426222305 | JHI_St_60k_v1 | DMT400065696 | GCTGGTTGGGATGGAAAGATTCGGACATTCCATAACTATGGATTGCCTGTCCGAGTTTAA |
All Microarray Probes Designed to Gene DMG400025559
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23278_PI426222305 | JHI_St_60k_v1 | DMT400065696 | GCTGGTTGGGATGGAAAGATTCGGACATTCCATAACTATGGATTGCCTGTCCGAGTTTAA |
| CUST_23409_PI426222305 | JHI_St_60k_v1 | DMT400065697 | CTCATCTGCAGATAGAAACTGCGATAGAATTGAATTCGATGAATCTTTACAGGACTAATT |
Microarray Signals from CUST_23278_PI426222305
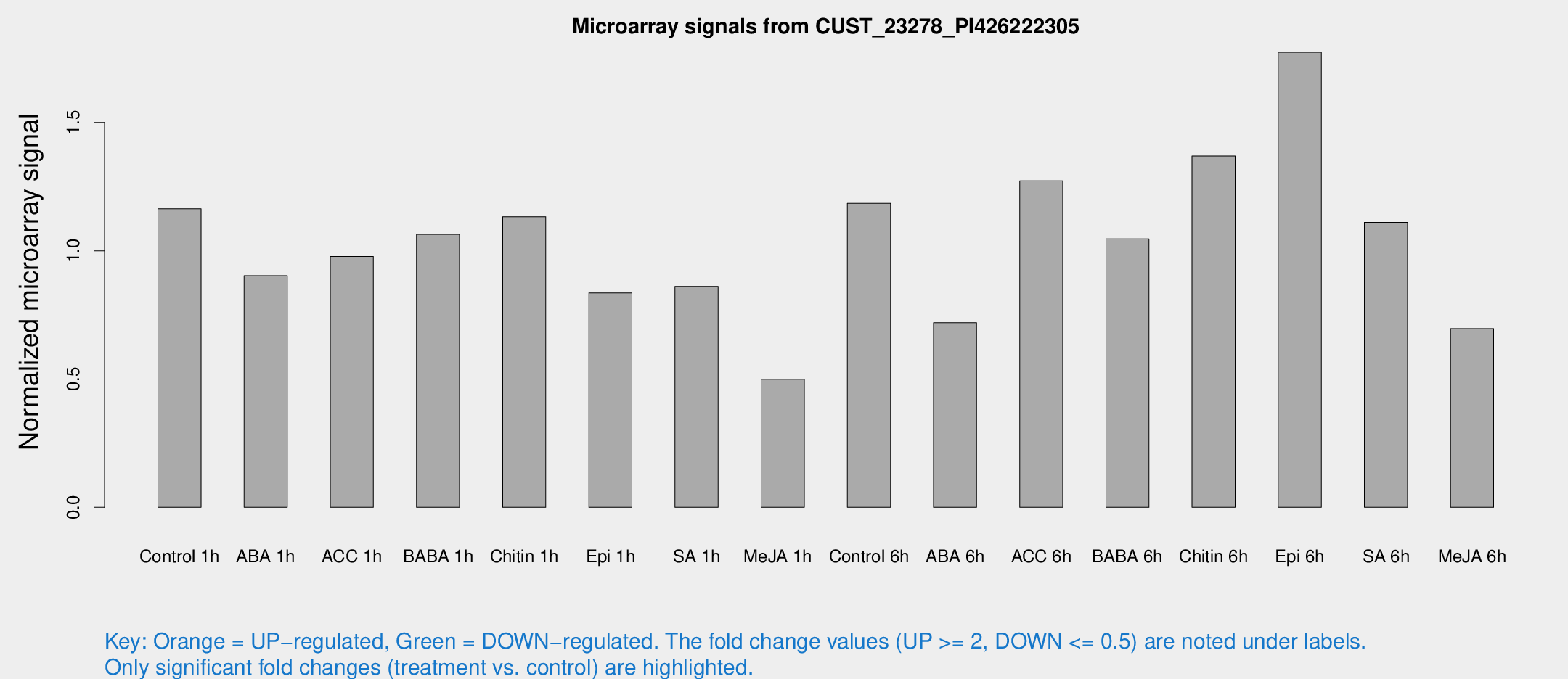
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 194.145 | 33.8837 | 1.16351 | 0.123999 |
| ABA 1h | 130.214 | 10.5359 | 0.902789 | 0.060437 |
| ACC 1h | 182.019 | 53.5703 | 0.977757 | 0.288011 |
| BABA 1h | 172.371 | 31.4865 | 1.0643 | 0.106753 |
| Chitin 1h | 174.021 | 36.2394 | 1.13282 | 0.206029 |
| Epi 1h | 118.171 | 11.3523 | 0.835705 | 0.0560041 |
| SA 1h | 148.992 | 29.4583 | 0.861318 | 0.155521 |
| Me-JA 1h | 67.7871 | 10.3932 | 0.499539 | 0.0429887 |
| Control 6h | 236.049 | 83.223 | 1.18484 | 0.556524 |
| ABA 6h | 130.242 | 28.0556 | 0.719779 | 0.190664 |
| ACC 6h | 250.813 | 57.994 | 1.27252 | 0.18973 |
| BABA 6h | 193.563 | 27.9976 | 1.04611 | 0.130851 |
| Chitin 6h | 236.685 | 14.2166 | 1.36939 | 0.0821153 |
| Epi 6h | 341.571 | 82.6267 | 1.7738 | 0.41688 |
| SA 6h | 185.902 | 39.1977 | 1.11103 | 0.144856 |
| Me-JA 6h | 119.455 | 26.7647 | 0.696737 | 0.138163 |
Source Transcript PGSC0003DMT400065696 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G48870.1 | +1 | 2e-118 | 369 | 242/657 (37%) | Transducin/WD40 repeat-like superfamily protein | chr1:18072325-18074457 REVERSE LENGTH=593 |
