Probe CUST_23105_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23105_PI426222305 | JHI_St_60k_v1 | DMT400021087 | GTTGTTCGATAAGGTCCACAAGTTAATTCGAACATCATCGTCAGATTGGAAAATGAATTA |
All Microarray Probes Designed to Gene DMG401008167
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22886_PI426222305 | JHI_St_60k_v1 | DMT400021089 | TGTAAGGAAGGCATCAACATCACCATTCTTGCTAAAAACATACATGTTGGTGGAGGATCC |
| CUST_23105_PI426222305 | JHI_St_60k_v1 | DMT400021087 | GTTGTTCGATAAGGTCCACAAGTTAATTCGAACATCATCGTCAGATTGGAAAATGAATTA |
Microarray Signals from CUST_23105_PI426222305
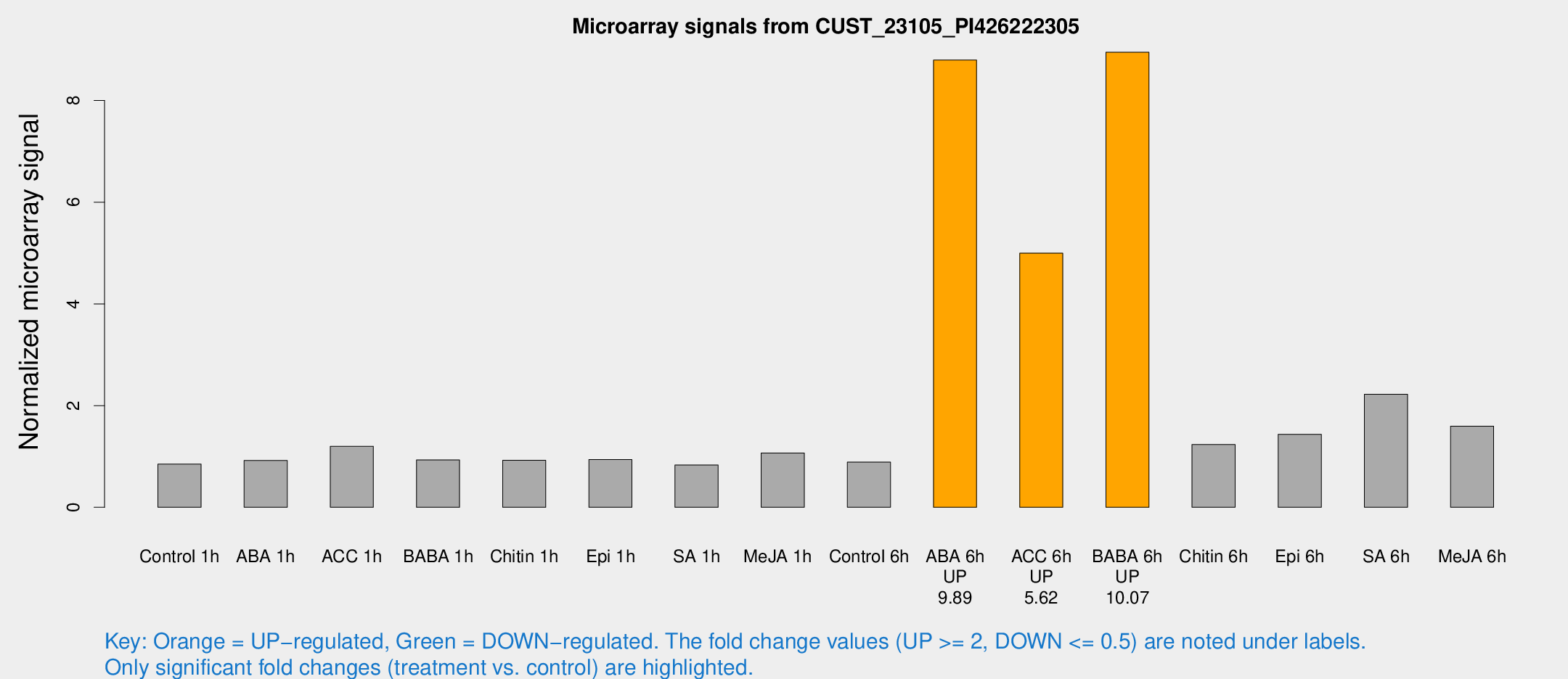
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.26172 | 3.6358 | 0.848314 | 0.491466 |
| ABA 1h | 6.00299 | 3.47997 | 0.921118 | 0.533792 |
| ACC 1h | 9.37282 | 4.66374 | 1.19822 | 0.60271 |
| BABA 1h | 6.631 | 3.8529 | 0.931008 | 0.539106 |
| Chitin 1h | 6.17223 | 3.60058 | 0.925157 | 0.535747 |
| Epi 1h | 6.01455 | 3.49817 | 0.939205 | 0.543866 |
| SA 1h | 6.29509 | 3.651 | 0.830834 | 0.481123 |
| Me-JA 1h | 6.45439 | 3.7658 | 1.06679 | 0.618491 |
| Control 6h | 6.57273 | 3.81658 | 0.888951 | 0.5154 |
| ABA 6h | 81.7013 | 29.8751 | 8.79503 | 3.812 |
| ACC 6h | 42.2475 | 5.01482 | 4.99816 | 0.587187 |
| BABA 6h | 75.6147 | 11.4149 | 8.94947 | 1.46117 |
| Chitin 6h | 10.7987 | 4.24963 | 1.23574 | 0.589961 |
| Epi 6h | 13.9675 | 5.70507 | 1.43496 | 0.763365 |
| SA 6h | 21.7063 | 11.4519 | 2.22302 | 1.53864 |
| Me-JA 6h | 12.8067 | 3.86282 | 1.59691 | 0.596404 |
Source Transcript PGSC0003DMT400021087 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G41690.1 | +2 | 2e-14 | 72 | 49/115 (43%) | heat shock transcription factor B3 | chr2:17381723-17382577 FORWARD LENGTH=244 |
