Probe CUST_23059_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23059_PI426222305 | JHI_St_60k_v1 | DMT400088273 | AGACTCGTGAAAAGGGTTCCTCTTATGTCCAATCAAATCCAAATGCGTGAACAATGCTAA |
All Microarray Probes Designed to Gene DMG400037844
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23059_PI426222305 | JHI_St_60k_v1 | DMT400088273 | AGACTCGTGAAAAGGGTTCCTCTTATGTCCAATCAAATCCAAATGCGTGAACAATGCTAA |
Microarray Signals from CUST_23059_PI426222305
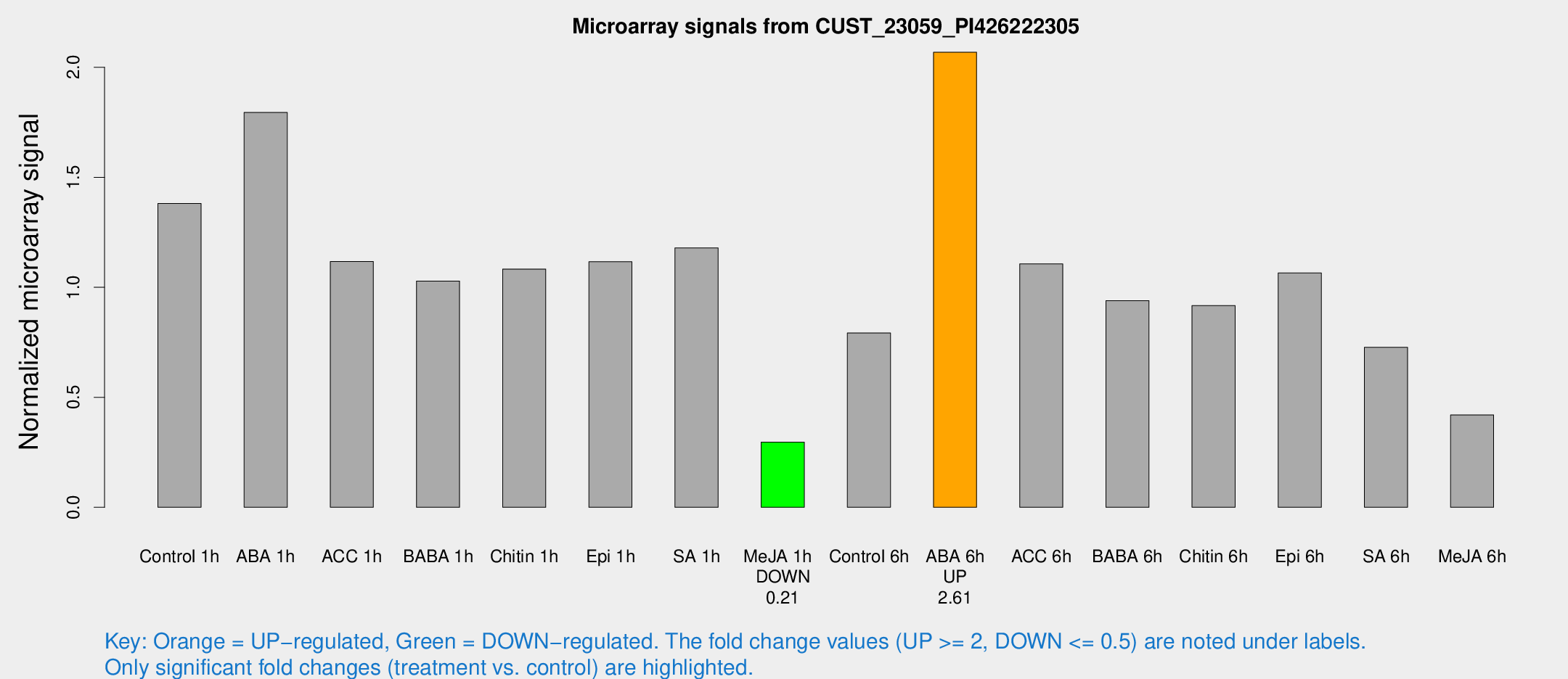
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 357.942 | 47.4495 | 1.38105 | 0.102332 |
| ABA 1h | 404.965 | 23.5798 | 1.79473 | 0.112691 |
| ACC 1h | 304.965 | 57.0272 | 1.11697 | 0.15185 |
| BABA 1h | 271.723 | 66.2453 | 1.02862 | 0.187913 |
| Chitin 1h | 251.218 | 26.744 | 1.08299 | 0.116896 |
| Epi 1h | 250.797 | 32.0109 | 1.11608 | 0.1371 |
| SA 1h | 324.875 | 75.3585 | 1.17892 | 0.225439 |
| Me-JA 1h | 63.6514 | 11.6989 | 0.296346 | 0.0357813 |
| Control 6h | 210.8 | 40.2224 | 0.792093 | 0.101948 |
| ABA 6h | 563.817 | 42.483 | 2.06862 | 0.120054 |
| ACC 6h | 323.69 | 19.0725 | 1.10644 | 0.166171 |
| BABA 6h | 270.129 | 22.3634 | 0.939572 | 0.0556849 |
| Chitin 6h | 249.019 | 14.8082 | 0.917168 | 0.0545242 |
| Epi 6h | 307.999 | 24.1598 | 1.06504 | 0.120411 |
| SA 6h | 187.438 | 27.721 | 0.72707 | 0.0542592 |
| Me-JA 6h | 113.967 | 28.3663 | 0.419746 | 0.0767951 |
Source Transcript PGSC0003DMT400088273 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G23180.1 | +1 | 1e-154 | 456 | 242/516 (47%) | cytochrome P450, family 96, subfamily A, polypeptide 1 | chr2:9874953-9876503 FORWARD LENGTH=516 |
