Probe CUST_23039_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23039_PI426222305 | JHI_St_60k_v1 | DMT400090220 | ATGAAACATGGTCTGAAAGTTAGGCTTGTCAAAAGGGTTCCTCTTATGTCCAATGCATGA |
All Microarray Probes Designed to Gene DMG400039791
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23039_PI426222305 | JHI_St_60k_v1 | DMT400090220 | ATGAAACATGGTCTGAAAGTTAGGCTTGTCAAAAGGGTTCCTCTTATGTCCAATGCATGA |
Microarray Signals from CUST_23039_PI426222305
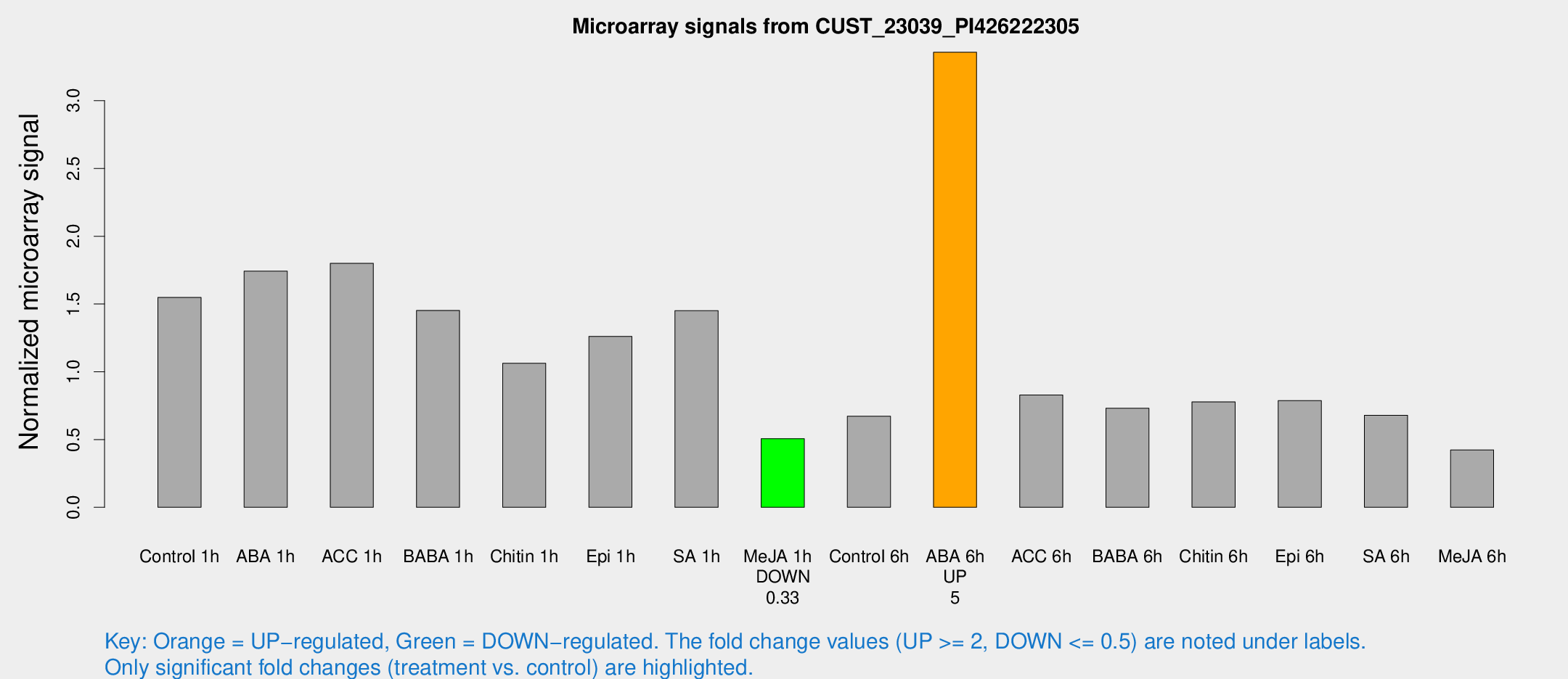
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 540.088 | 56.1478 | 1.54787 | 0.0897592 |
| ABA 1h | 542.691 | 73.1247 | 1.74213 | 0.174966 |
| ACC 1h | 749.319 | 278.57 | 1.79966 | 0.674349 |
| BABA 1h | 503.849 | 98.3939 | 1.45226 | 0.166917 |
| Chitin 1h | 335.541 | 45.4499 | 1.06276 | 0.0622048 |
| Epi 1h | 380.514 | 37.1341 | 1.26107 | 0.115381 |
| SA 1h | 528.868 | 89.8046 | 1.4506 | 0.154144 |
| Me-JA 1h | 143.024 | 8.78651 | 0.50675 | 0.0374802 |
| Control 6h | 247.802 | 57.2105 | 0.672121 | 0.122123 |
| ABA 6h | 1233.5 | 71.389 | 3.3575 | 0.194031 |
| ACC 6h | 329.431 | 23.4266 | 0.828133 | 0.134785 |
| BABA 6h | 288.022 | 40.5769 | 0.730862 | 0.0755423 |
| Chitin 6h | 292.552 | 44.703 | 0.777604 | 0.140102 |
| Epi 6h | 310.647 | 37.1012 | 0.787391 | 0.12541 |
| SA 6h | 237.086 | 34.6757 | 0.678358 | 0.0403928 |
| Me-JA 6h | 164.054 | 58.1383 | 0.423598 | 0.116203 |
Source Transcript PGSC0003DMT400090220 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G39490.1 | +1 | 6e-160 | 468 | 238/488 (49%) | cytochrome P450, family 96, subfamily A, polypeptide 10 | chr4:18365229-18366788 FORWARD LENGTH=519 |
