Probe CUST_22893_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22893_PI426222305 | JHI_St_60k_v1 | DMT400060940 | GCAGCACATCGGTGACAAATCAAAGAGAAAGTTGTAAGGTTTTAAGCATAGTAGTTTTTT |
All Microarray Probes Designed to Gene DMG400023703
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22893_PI426222305 | JHI_St_60k_v1 | DMT400060940 | GCAGCACATCGGTGACAAATCAAAGAGAAAGTTGTAAGGTTTTAAGCATAGTAGTTTTTT |
Microarray Signals from CUST_22893_PI426222305
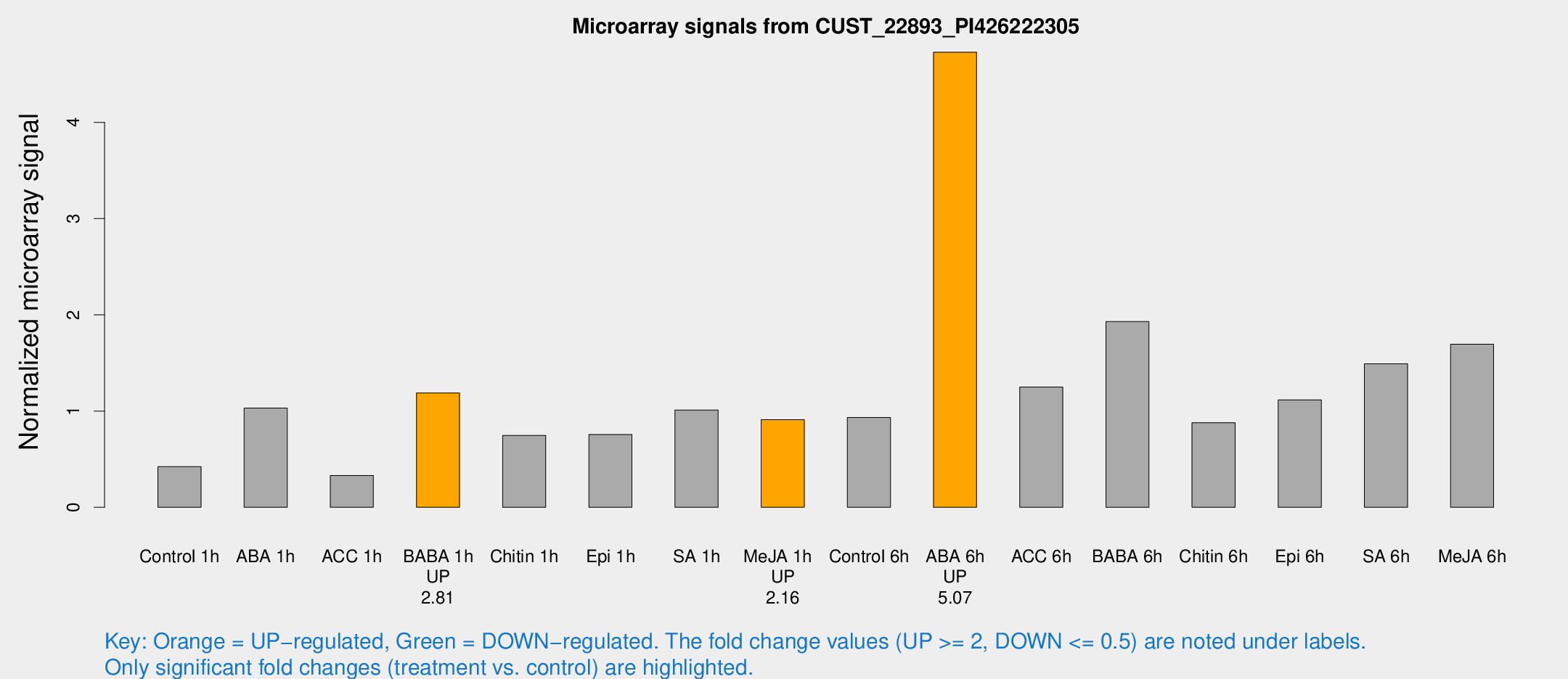
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 279.986 | 16.5695 | 0.422362 | 0.0271058 |
| ABA 1h | 718.053 | 299.971 | 1.03208 | 0.387195 |
| ACC 1h | 229.786 | 35.4818 | 0.330128 | 0.0295079 |
| BABA 1h | 806.949 | 187.705 | 1.18845 | 0.192707 |
| Chitin 1h | 458.833 | 85.3935 | 0.746875 | 0.0875957 |
| Epi 1h | 462.055 | 108.953 | 0.755557 | 0.193952 |
| SA 1h | 747.905 | 221.014 | 1.01155 | 0.267738 |
| Me-JA 1h | 496.385 | 44.0893 | 0.911525 | 0.0530571 |
| Control 6h | 686.372 | 198.191 | 0.933426 | 0.248055 |
| ABA 6h | 3430.17 | 621.704 | 4.72846 | 0.569033 |
| ACC 6h | 959.194 | 100.367 | 1.24954 | 0.0723671 |
| BABA 6h | 1526.63 | 410.816 | 1.92999 | 0.43625 |
| Chitin 6h | 619.796 | 36.0524 | 0.879442 | 0.0511351 |
| Epi 6h | 848.109 | 107.085 | 1.1166 | 0.0828345 |
| SA 6h | 1049.63 | 249.865 | 1.4913 | 0.250725 |
| Me-JA 6h | 1198.34 | 285.737 | 1.69409 | 0.373061 |
Source Transcript PGSC0003DMT400060940 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G15670.1 | +3 | 3e-66 | 219 | 128/363 (35%) | Galactose oxidase/kelch repeat superfamily protein | chr1:5390119-5391198 FORWARD LENGTH=359 |
