Probe CUST_22771_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22771_PI426222305 | JHI_St_60k_v1 | DMT400078033 | GAGACTATCAACTAAGAGAGTTGTGGATGTATACGGTAATACAACATTGAATGTCTTTCA |
All Microarray Probes Designed to Gene DMG400030351
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22771_PI426222305 | JHI_St_60k_v1 | DMT400078033 | GAGACTATCAACTAAGAGAGTTGTGGATGTATACGGTAATACAACATTGAATGTCTTTCA |
Microarray Signals from CUST_22771_PI426222305
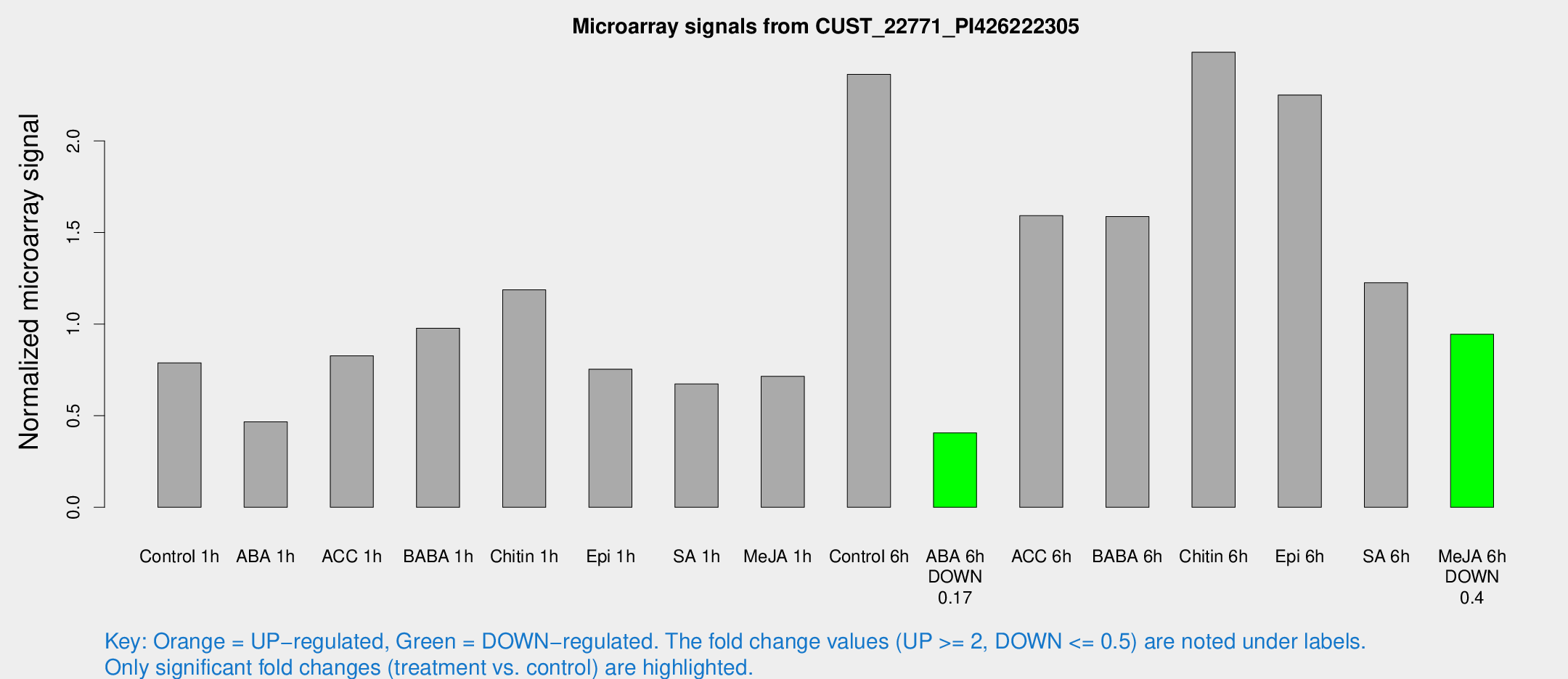
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 189.483 | 34.2458 | 0.78883 | 0.0969159 |
| ABA 1h | 96.7986 | 7.51093 | 0.466551 | 0.0431001 |
| ACC 1h | 204.9 | 36.0125 | 0.826923 | 0.100653 |
| BABA 1h | 229.067 | 44.1292 | 0.977786 | 0.109575 |
| Chitin 1h | 252.528 | 29.9967 | 1.18682 | 0.105604 |
| Epi 1h | 152.177 | 9.4108 | 0.753864 | 0.0466258 |
| SA 1h | 168.645 | 33.8284 | 0.673325 | 0.107876 |
| Me-JA 1h | 138.107 | 16.2416 | 0.714836 | 0.0501398 |
| Control 6h | 574.109 | 106.304 | 2.36308 | 0.258214 |
| ABA 6h | 102.235 | 12.1133 | 0.406375 | 0.050485 |
| ACC 6h | 450.271 | 99.1037 | 1.59176 | 0.256277 |
| BABA 6h | 438.28 | 105.701 | 1.58727 | 0.410939 |
| Chitin 6h | 616.589 | 35.8231 | 2.48417 | 0.144272 |
| Epi 6h | 596.008 | 51.7484 | 2.2509 | 0.311487 |
| SA 6h | 284.637 | 23.4263 | 1.22587 | 0.0726511 |
| Me-JA 6h | 236.564 | 65.535 | 0.944675 | 0.18638 |
Source Transcript PGSC0003DMT400078033 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G07270.1 | +2 | 3e-175 | 514 | 281/460 (61%) | Cell division control, Cdc6 | chr1:2229757-2232897 REVERSE LENGTH=505 |
