Probe CUST_22607_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22607_PI426222305 | JHI_St_60k_v1 | DMT400078120 | GGGGTTGTCTTTGTTTACTAGGAACTTGTCTTGCTTTTATTCTTCATAGAAGAACACAAA |
All Microarray Probes Designed to Gene DMG400030381
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22607_PI426222305 | JHI_St_60k_v1 | DMT400078120 | GGGGTTGTCTTTGTTTACTAGGAACTTGTCTTGCTTTTATTCTTCATAGAAGAACACAAA |
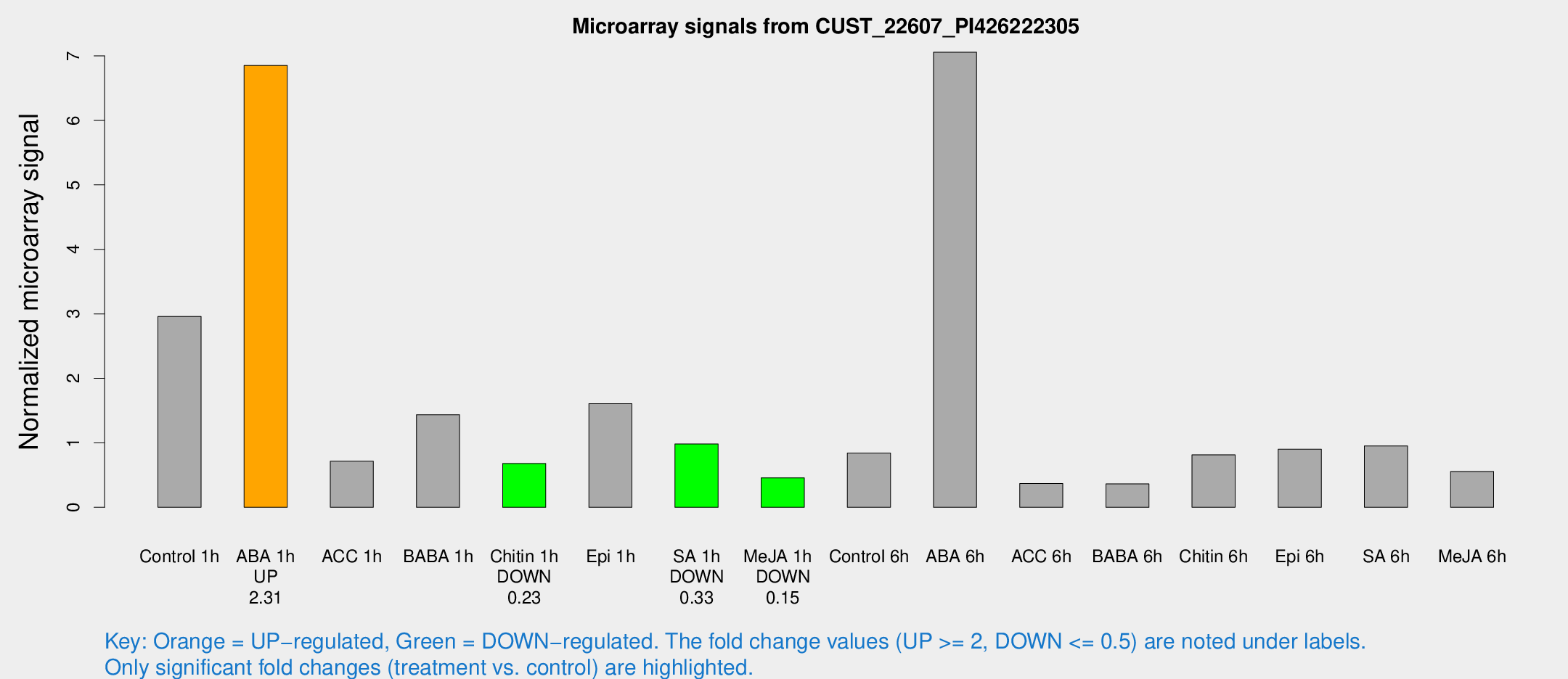
Microarray Signals from CUST_22607_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 141.348 | 19.2209 | 2.96042 | 0.290602 |
| ABA 1h | 285.952 | 25.9648 | 6.85167 | 1.1339 |
| ACC 1h | 51.0255 | 22.9162 | 0.715577 | 0.672981 |
| BABA 1h | 81.7139 | 35.8982 | 1.43569 | 0.679999 |
| Chitin 1h | 29.0441 | 4.00829 | 0.679481 | 0.0955483 |
| Epi 1h | 66.602 | 9.91755 | 1.60746 | 0.24388 |
| SA 1h | 47.448 | 4.57977 | 0.982244 | 0.0949985 |
| Me-JA 1h | 17.6673 | 3.84239 | 0.457334 | 0.104519 |
| Control 6h | 53.625 | 21.939 | 0.842151 | 0.58042 |
| ABA 6h | 351.445 | 20.7198 | 7.05664 | 0.461219 |
| ACC 6h | 21.7056 | 6.95046 | 0.368589 | 0.088914 |
| BABA 6h | 21.1637 | 6.53268 | 0.362967 | 0.143273 |
| Chitin 6h | 43.6697 | 11.1645 | 0.814468 | 0.22606 |
| Epi 6h | 49.7181 | 10.2394 | 0.900245 | 0.283755 |
| SA 6h | 44.4266 | 4.78512 | 0.952668 | 0.108305 |
| Me-JA 6h | 34.0934 | 18.3075 | 0.555433 | 0.297763 |
Source Transcript PGSC0003DMT400078120 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G30300.1 | +3 | 9e-111 | 343 | 260/516 (50%) | Major facilitator superfamily protein | chr2:12919401-12921222 FORWARD LENGTH=500 |
