Probe CUST_22402_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22402_PI426222305 | JHI_St_60k_v1 | DMT400039412 | GCGACTGGAATAAAAAATTGAGTATATATGACTGTATAGCGAGCTTCAAGATAACTTGTG |
All Microarray Probes Designed to Gene DMG400015242
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22402_PI426222305 | JHI_St_60k_v1 | DMT400039412 | GCGACTGGAATAAAAAATTGAGTATATATGACTGTATAGCGAGCTTCAAGATAACTTGTG |
Microarray Signals from CUST_22402_PI426222305
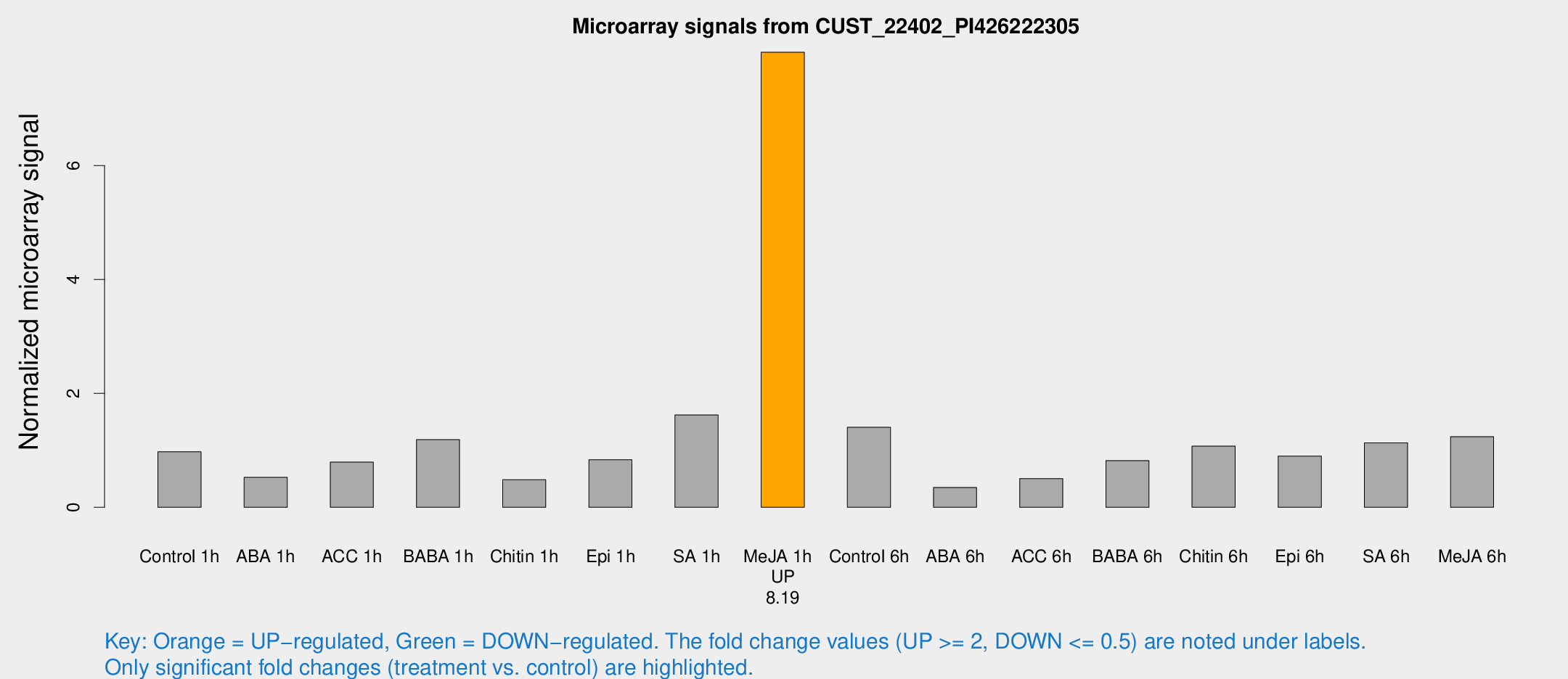
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 85.7571 | 23.4618 | 0.976194 | 0.193865 |
| ABA 1h | 41.3399 | 10.6325 | 0.525738 | 0.157998 |
| ACC 1h | 70.6478 | 15.7421 | 0.792815 | 0.134354 |
| BABA 1h | 96.057 | 13.6057 | 1.18768 | 0.080518 |
| Chitin 1h | 35.6889 | 3.69791 | 0.483518 | 0.0501635 |
| Epi 1h | 60.3394 | 8.21281 | 0.833913 | 0.0948959 |
| SA 1h | 139.159 | 19.0837 | 1.62091 | 0.180359 |
| Me-JA 1h | 548.268 | 84.1802 | 7.99148 | 0.544015 |
| Control 6h | 123.709 | 34.065 | 1.404 | 0.286512 |
| ABA 6h | 30.9947 | 3.99834 | 0.349406 | 0.0663771 |
| ACC 6h | 49.6064 | 10.5657 | 0.503439 | 0.0674627 |
| BABA 6h | 78.319 | 14.2681 | 0.81998 | 0.205138 |
| Chitin 6h | 95.4365 | 11.7105 | 1.07464 | 0.112929 |
| Epi 6h | 90.0139 | 22.5167 | 0.900284 | 0.345389 |
| SA 6h | 95.4651 | 17.8913 | 1.13235 | 0.113356 |
| Me-JA 6h | 105.009 | 18.2017 | 1.24097 | 0.158886 |
Source Transcript PGSC0003DMT400039412 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G48520.1 | +1 | 0.0 | 576 | 286/468 (61%) | cytochrome P450, family 94, subfamily B, polypeptide 3 | chr3:17975104-17976624 REVERSE LENGTH=506 |
