Probe CUST_22393_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22393_PI426222305 | JHI_St_60k_v1 | DMT400039272 | CGAGATTAAACCGGTGGTTAATGTAGAAAAACCGGTTTTTGTTCCTCTTTTGACTGCACA |
All Microarray Probes Designed to Gene DMG400015185
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22393_PI426222305 | JHI_St_60k_v1 | DMT400039272 | CGAGATTAAACCGGTGGTTAATGTAGAAAAACCGGTTTTTGTTCCTCTTTTGACTGCACA |
Microarray Signals from CUST_22393_PI426222305
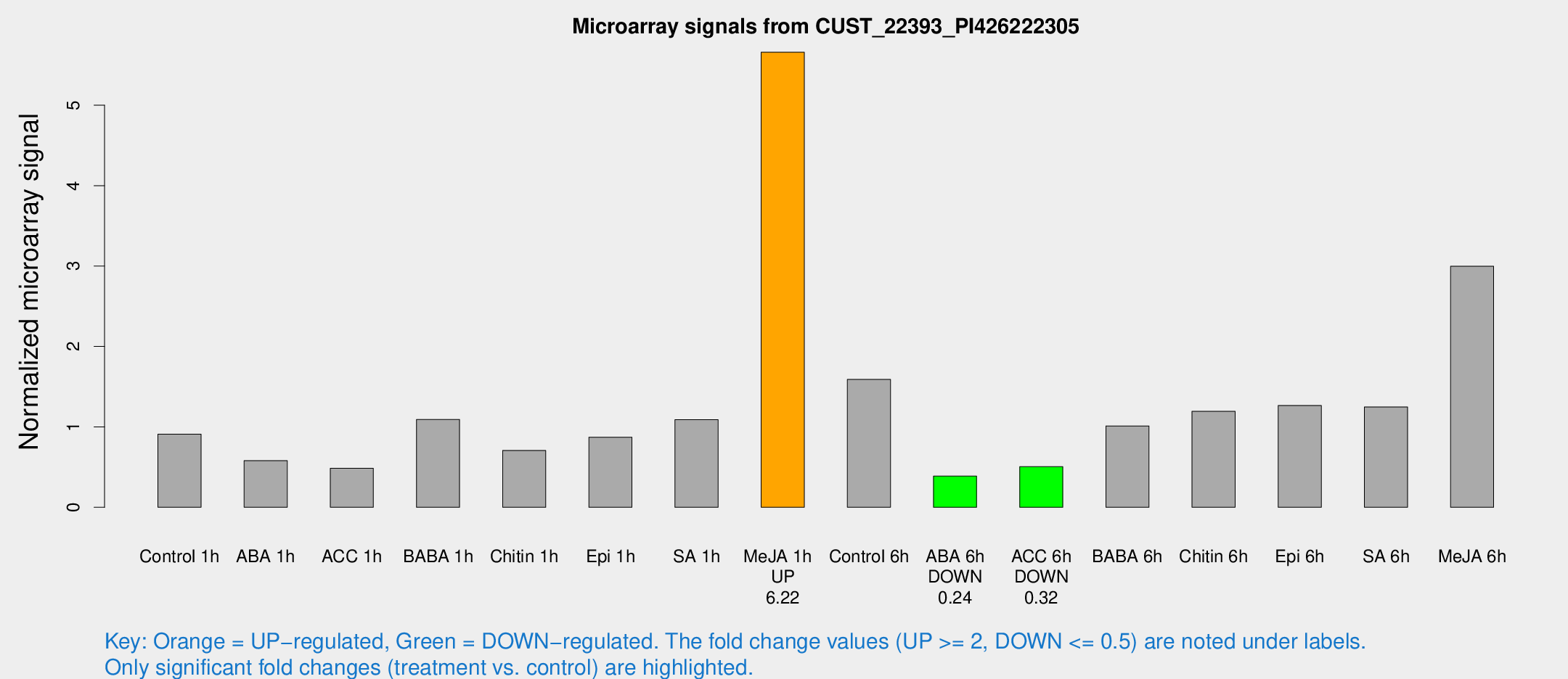
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 96.2171 | 15.0543 | 0.910478 | 0.0888938 |
| ABA 1h | 55.3322 | 12.3927 | 0.58009 | 0.0856894 |
| ACC 1h | 53.7598 | 10.6905 | 0.484564 | 0.0731074 |
| BABA 1h | 112.385 | 19.6328 | 1.09241 | 0.0994704 |
| Chitin 1h | 66.5175 | 8.20516 | 0.706959 | 0.0721544 |
| Epi 1h | 78.5562 | 7.7735 | 0.872339 | 0.0925664 |
| SA 1h | 120.122 | 23.2649 | 1.08979 | 0.191804 |
| Me-JA 1h | 485.555 | 66.2097 | 5.65906 | 0.375367 |
| Control 6h | 172.704 | 38.3209 | 1.59032 | 0.227158 |
| ABA 6h | 42.6338 | 4.45884 | 0.38804 | 0.0410269 |
| ACC 6h | 62.3939 | 13.4294 | 0.505478 | 0.0510156 |
| BABA 6h | 123.359 | 29.2972 | 1.0118 | 0.263312 |
| Chitin 6h | 133.442 | 18.045 | 1.19308 | 0.149863 |
| Epi 6h | 153.547 | 30.6512 | 1.26613 | 0.34902 |
| SA 6h | 131.665 | 23.7625 | 1.24741 | 0.11462 |
| Me-JA 6h | 307.452 | 18.1101 | 2.99745 | 0.234846 |
Source Transcript PGSC0003DMT400039272 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G48520.1 | +3 | 0.0 | 590 | 296/471 (63%) | cytochrome P450, family 94, subfamily B, polypeptide 3 | chr3:17975104-17976624 REVERSE LENGTH=506 |
