Probe CUST_22210_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22210_PI426222305 | JHI_St_60k_v1 | DMT400023467 | TGGTTTTCATCTGTGAAGAAAGTTTTTAAACAGTCTCCTAAAGATTCACCAGACAAGAAG |
All Microarray Probes Designed to Gene DMG400009086
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22193_PI426222305 | JHI_St_60k_v1 | DMT400023468 | GGCGATGATCGGGCTTCACCATTAGGACCACATCCCTGGAGATATAATTTTAGTTGATTT |
| CUST_22210_PI426222305 | JHI_St_60k_v1 | DMT400023467 | TGGTTTTCATCTGTGAAGAAAGTTTTTAAACAGTCTCCTAAAGATTCACCAGACAAGAAG |
Microarray Signals from CUST_22210_PI426222305
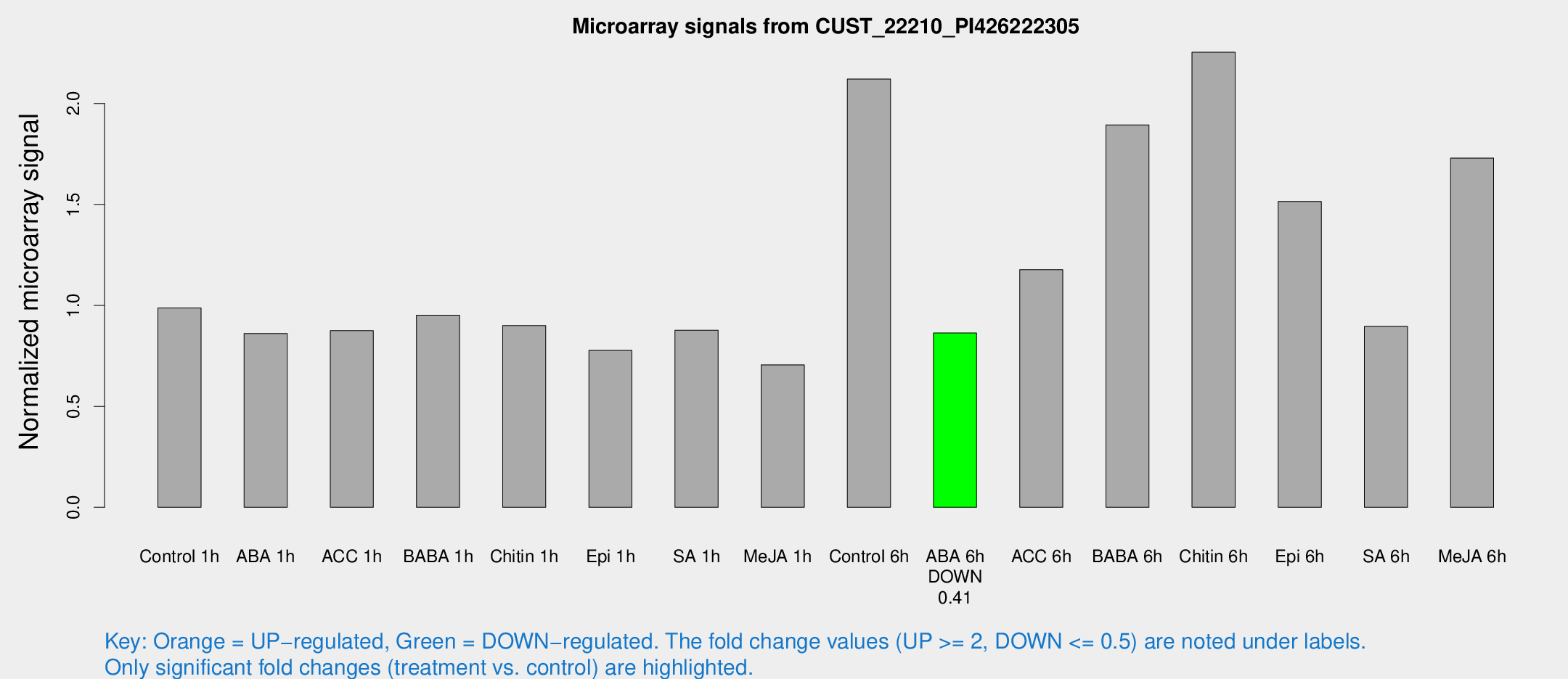
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 58.4345 | 9.77211 | 0.98707 | 0.106074 |
| ABA 1h | 44.7437 | 6.79526 | 0.860511 | 0.0942068 |
| ACC 1h | 56.341 | 16.9992 | 0.874994 | 0.20821 |
| BABA 1h | 53.9345 | 8.0586 | 0.951911 | 0.0893833 |
| Chitin 1h | 47.105 | 5.0286 | 0.899929 | 0.134973 |
| Epi 1h | 39.2617 | 4.9154 | 0.777273 | 0.0845922 |
| SA 1h | 53.1998 | 8.1915 | 0.877188 | 0.14998 |
| Me-JA 1h | 33.9341 | 5.49381 | 0.705572 | 0.130941 |
| Control 6h | 126.809 | 22.395 | 2.12122 | 0.227651 |
| ABA 6h | 53.0822 | 4.98447 | 0.863155 | 0.0813599 |
| ACC 6h | 85.6293 | 26.1104 | 1.17663 | 0.239451 |
| BABA 6h | 126.702 | 26.0339 | 1.89402 | 0.365432 |
| Chitin 6h | 138.131 | 9.02977 | 2.25405 | 0.147388 |
| Epi 6h | 112.495 | 36.288 | 1.51506 | 0.787314 |
| SA 6h | 70.6265 | 29.4595 | 0.896135 | 0.997565 |
| Me-JA 6h | 110.844 | 32.0713 | 1.72965 | 0.508838 |
Source Transcript PGSC0003DMT400023467 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G49260.3 | +1 | 1e-53 | 192 | 212/473 (45%) | IQ-domain 21 | chr3:18262755-18265859 FORWARD LENGTH=472 |
