Probe CUST_21237_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21237_PI426222305 | JHI_St_60k_v1 | DMT400020334 | CTAGTCATTACCTAAGGAGTTGGAGACCTACAACCACTCAACCATGTAACAATATGTAAA |
All Microarray Probes Designed to Gene DMG400007862
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21237_PI426222305 | JHI_St_60k_v1 | DMT400020334 | CTAGTCATTACCTAAGGAGTTGGAGACCTACAACCACTCAACCATGTAACAATATGTAAA |
| CUST_21261_PI426222305 | JHI_St_60k_v1 | DMT400020333 | CTAGTCATTACCTAAGGAGTTGGAGACCTACAACCACTCAACCATGTAACAATATGTAAA |
Microarray Signals from CUST_21237_PI426222305
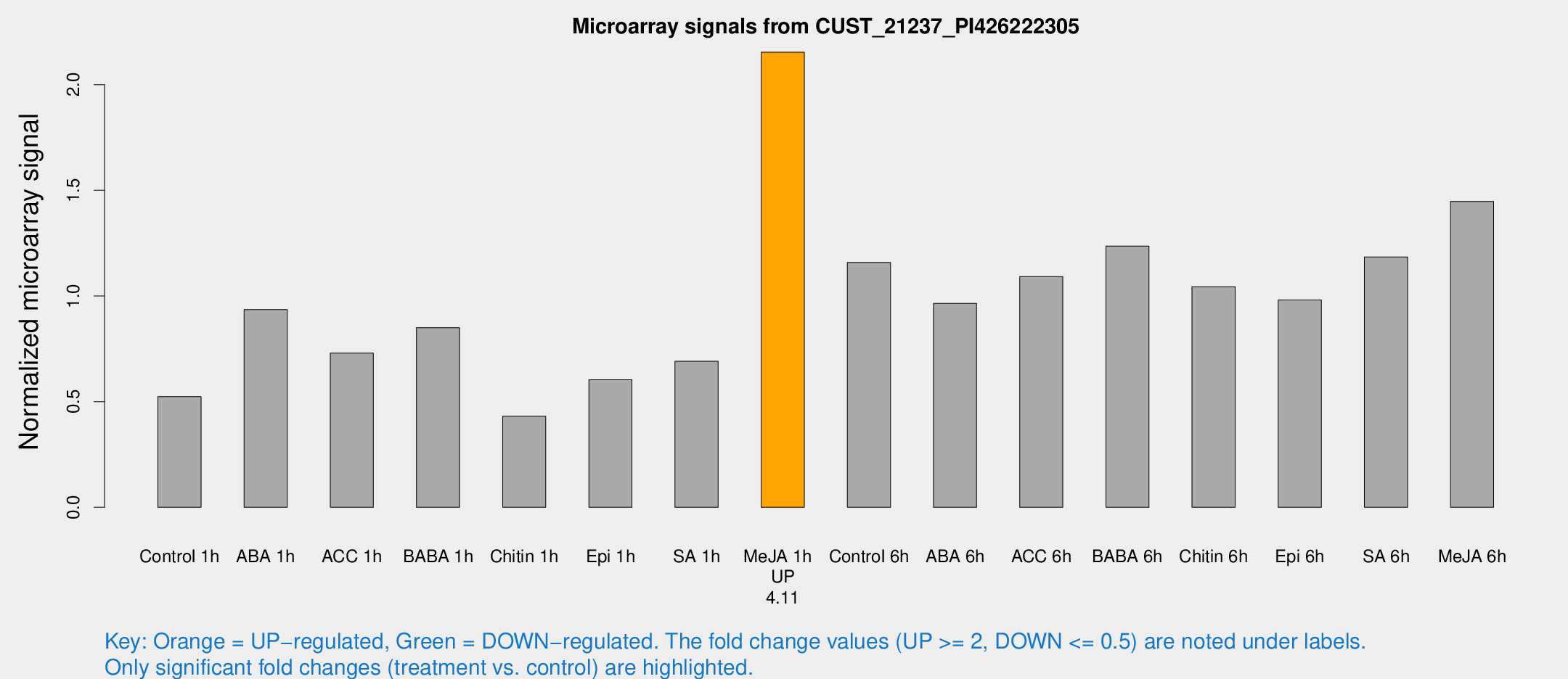
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.402 | 3.12878 | 0.523983 | 0.184645 |
| ABA 1h | 15.7249 | 4.63889 | 0.935347 | 0.272534 |
| ACC 1h | 14.7675 | 4.49311 | 0.729563 | 0.248364 |
| BABA 1h | 17.2826 | 6.13543 | 0.849928 | 0.355063 |
| Chitin 1h | 6.8815 | 3.09268 | 0.431279 | 0.204071 |
| Epi 1h | 9.2322 | 3.09731 | 0.603897 | 0.211603 |
| SA 1h | 15.5256 | 7.02956 | 0.690743 | 0.445803 |
| Me-JA 1h | 31.1514 | 3.89154 | 2.15319 | 0.255728 |
| Control 6h | 27.6911 | 11.1357 | 1.15851 | 0.81868 |
| ABA 6h | 19.2758 | 4.86642 | 0.965151 | 0.254766 |
| ACC 6h | 21.8799 | 4.01965 | 1.09188 | 0.200473 |
| BABA 6h | 27.5034 | 10.0093 | 1.23603 | 0.429939 |
| Chitin 6h | 20.5555 | 5.21674 | 1.04338 | 0.223454 |
| Epi 6h | 23.6659 | 8.8351 | 0.981115 | 0.685987 |
| SA 6h | 21.1674 | 4.00469 | 1.18428 | 0.215501 |
| Me-JA 6h | 26.6597 | 5.78767 | 1.44728 | 0.268591 |
Source Transcript PGSC0003DMT400020334 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G03390.1 | +3 | 0.0 | 815 | 474/783 (61%) | STRUBBELIG-receptor family 3 | chr4:1490912-1494553 REVERSE LENGTH=776 |
