Probe CUST_21190_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21190_PI426222305 | JHI_St_60k_v1 | DMT400020345 | CCTGCATGCTCATTCTTACTATAATGTTCTCTTGTTCCTTAATTATGGGACATCTAACAT |
All Microarray Probes Designed to Gene DMG400007869
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21190_PI426222305 | JHI_St_60k_v1 | DMT400020345 | CCTGCATGCTCATTCTTACTATAATGTTCTCTTGTTCCTTAATTATGGGACATCTAACAT |
| CUST_21247_PI426222305 | JHI_St_60k_v1 | DMT400020346 | ACAGATGAAGATGATGATGATGATGACTTTGAGAAAGAAGTTGCTTCTTGCCGCAATAAT |
Microarray Signals from CUST_21190_PI426222305
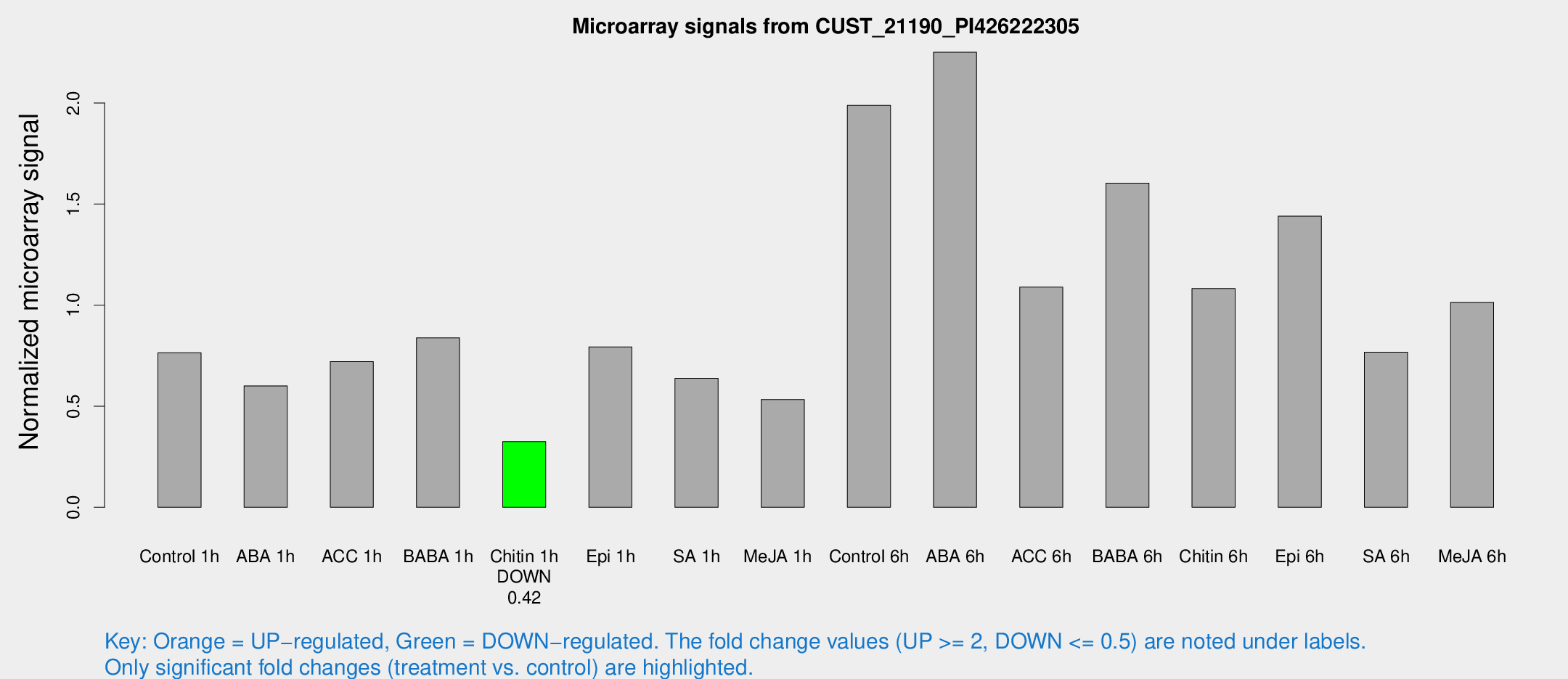
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 14.1141 | 3.16993 | 0.76478 | 0.172527 |
| ABA 1h | 11.1869 | 3.95756 | 0.600706 | 0.226148 |
| ACC 1h | 14.7552 | 3.93951 | 0.720968 | 0.246592 |
| BABA 1h | 16.18 | 4.54057 | 0.838332 | 0.213965 |
| Chitin 1h | 5.23122 | 3.05483 | 0.324644 | 0.182747 |
| Epi 1h | 14.5304 | 4.47982 | 0.793496 | 0.351587 |
| SA 1h | 14.172 | 5.30231 | 0.637939 | 0.368347 |
| Me-JA 1h | 10.1208 | 5.092 | 0.533499 | 0.247116 |
| Control 6h | 39.6001 | 9.86514 | 1.98787 | 0.420723 |
| ABA 6h | 47.7032 | 13.0588 | 2.25093 | 0.550718 |
| ACC 6h | 23.9889 | 5.22259 | 1.08927 | 0.182036 |
| BABA 6h | 40.2229 | 14.4551 | 1.60359 | 0.849074 |
| Chitin 6h | 23.4263 | 7.31758 | 1.08192 | 0.353059 |
| Epi 6h | 32.2446 | 8.38588 | 1.44041 | 0.523949 |
| SA 6h | 14.5879 | 3.47189 | 0.767016 | 0.208718 |
| Me-JA 6h | 25.0519 | 9.99942 | 1.0138 | 0.724622 |
Source Transcript PGSC0003DMT400020345 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G50180.1 | +3 | 9e-41 | 157 | 128/362 (35%) | NB-ARC domain-containing disease resistance protein | chr1:18584235-18587136 FORWARD LENGTH=857 |
