Probe CUST_21131_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21131_PI426222305 | JHI_St_60k_v1 | DMT400020318 | ATGAACATTTCATGATGTTCCTGCTTAGAATCCCAGACCTACGAGATGTACAGCTGAGAG |
All Microarray Probes Designed to Gene DMG400007856
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21131_PI426222305 | JHI_St_60k_v1 | DMT400020318 | ATGAACATTTCATGATGTTCCTGCTTAGAATCCCAGACCTACGAGATGTACAGCTGAGAG |
Microarray Signals from CUST_21131_PI426222305
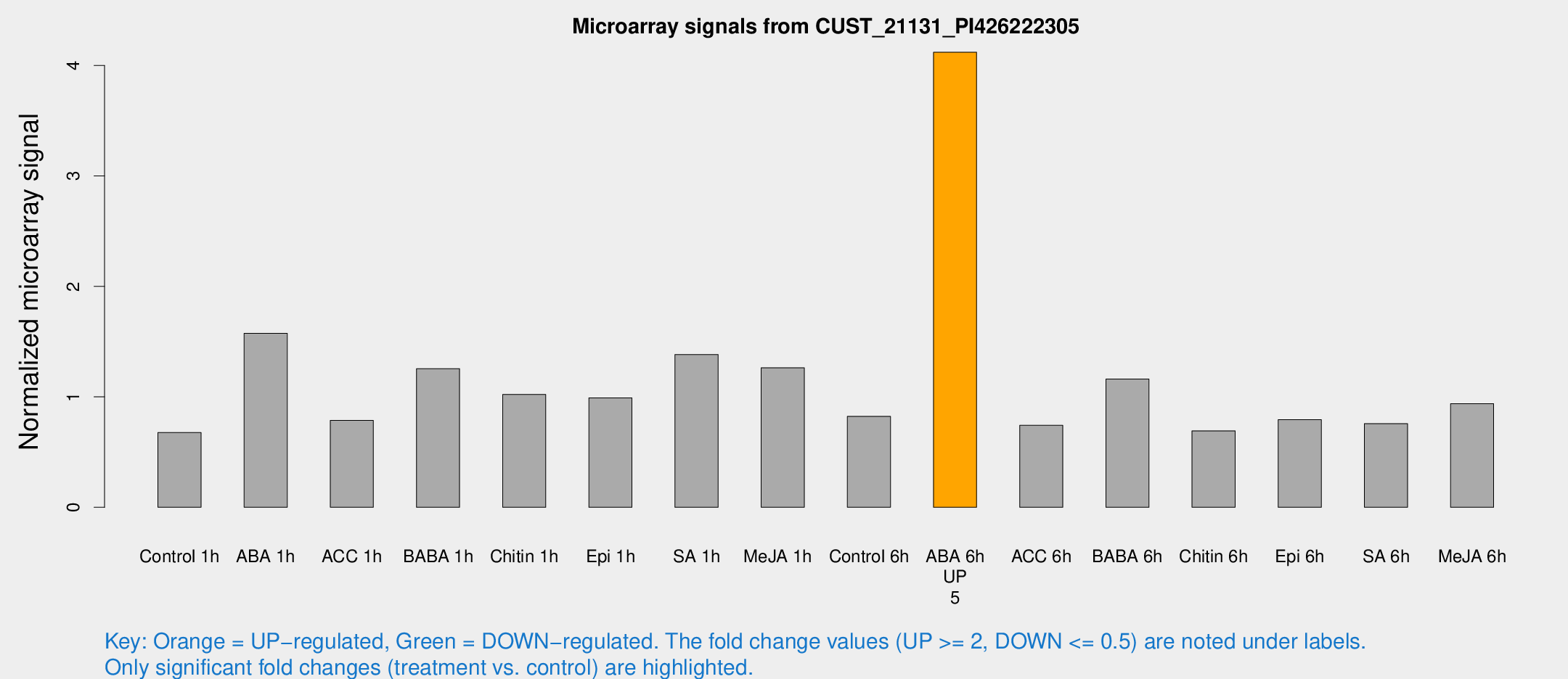
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 168.719 | 15.7919 | 0.677496 | 0.0426079 |
| ABA 1h | 390.231 | 143.711 | 1.57471 | 0.465656 |
| ACC 1h | 206.336 | 38.4035 | 0.786261 | 0.0952007 |
| BABA 1h | 310.705 | 61.6309 | 1.25512 | 0.155163 |
| Chitin 1h | 226.974 | 13.5572 | 1.02201 | 0.0899108 |
| Epi 1h | 218.287 | 35.0582 | 0.991349 | 0.166344 |
| SA 1h | 365.48 | 75.8835 | 1.38285 | 0.234596 |
| Me-JA 1h | 257.256 | 26.258 | 1.26277 | 0.1203 |
| Control 6h | 218.917 | 52.5908 | 0.82353 | 0.162551 |
| ABA 6h | 1099.95 | 135.218 | 4.12041 | 0.356767 |
| ACC 6h | 223.201 | 55.5533 | 0.742366 | 0.103612 |
| BABA 6h | 326.748 | 45.3978 | 1.16076 | 0.117507 |
| Chitin 6h | 183.972 | 18.1663 | 0.692616 | 0.0582243 |
| Epi 6h | 228.223 | 37.8647 | 0.794287 | 0.100637 |
| SA 6h | 192.85 | 39.9221 | 0.756893 | 0.0954185 |
| Me-JA 6h | 251.9 | 67.2278 | 0.937924 | 0.245185 |
Source Transcript PGSC0003DMT400020318 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G42620.1 | +1 | 0.0 | 775 | 417/728 (57%) | RNI-like superfamily protein | chr2:17756170-17758251 FORWARD LENGTH=693 |
