Probe CUST_21083_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21083_PI426222305 | JHI_St_60k_v1 | DMT400020228 | CCCTTTACTGAAAGGGCACACATTTAAAATTCAGCTTCTGGTAGCATAAACTTTAGATTA |
All Microarray Probes Designed to Gene DMG400007819
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21083_PI426222305 | JHI_St_60k_v1 | DMT400020228 | CCCTTTACTGAAAGGGCACACATTTAAAATTCAGCTTCTGGTAGCATAAACTTTAGATTA |
| CUST_21162_PI426222305 | JHI_St_60k_v1 | DMT400020227 | CCCTTTACTGAAAGGGCACACATTTAAAATTCAGCTTCTGGTAGCATAAACTTTAGATTA |
Microarray Signals from CUST_21083_PI426222305
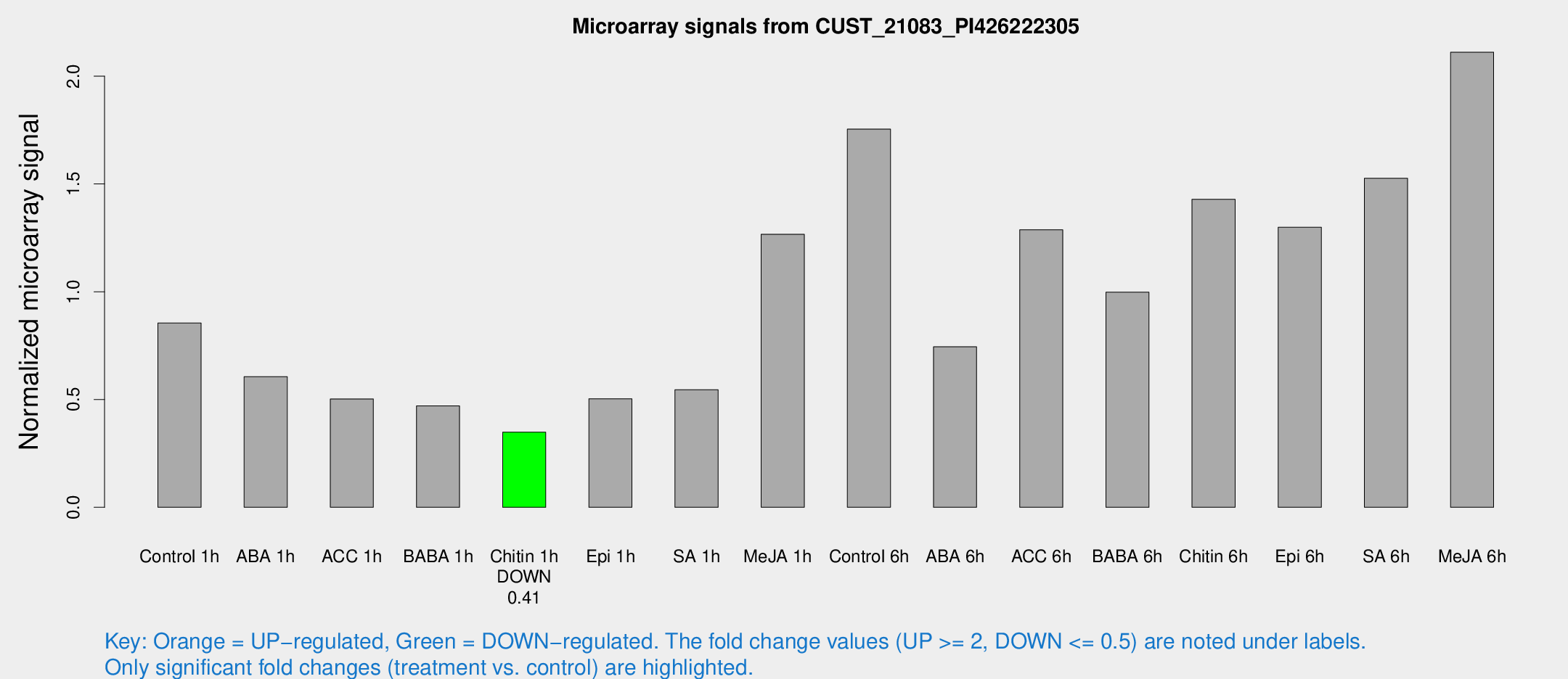
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 816.89 | 180.883 | 0.854829 | 0.13457 |
| ABA 1h | 495.466 | 51.809 | 0.606357 | 0.0614954 |
| ACC 1h | 576.244 | 232.978 | 0.502576 | 0.230747 |
| BABA 1h | 423.759 | 60.1588 | 0.470344 | 0.0275487 |
| Chitin 1h | 287.632 | 19.553 | 0.348364 | 0.0340382 |
| Epi 1h | 401.82 | 36.682 | 0.503546 | 0.0430014 |
| SA 1h | 565.652 | 187.809 | 0.545631 | 0.182363 |
| Me-JA 1h | 955.806 | 104.191 | 1.26661 | 0.073292 |
| Control 6h | 1668.66 | 318.868 | 1.75466 | 0.193567 |
| ABA 6h | 742.072 | 111.855 | 0.74462 | 0.0864277 |
| ACC 6h | 1355.13 | 86.7685 | 1.28756 | 0.0971887 |
| BABA 6h | 1033.77 | 119.224 | 0.99788 | 0.120252 |
| Chitin 6h | 1396.25 | 94.4611 | 1.42905 | 0.0826203 |
| Epi 6h | 1354.75 | 151.939 | 1.2991 | 0.187425 |
| SA 6h | 1388.23 | 105.097 | 1.52678 | 0.0882643 |
| Me-JA 6h | 1931.03 | 141.446 | 2.11119 | 0.121962 |
Source Transcript PGSC0003DMT400020228 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G25620.1 | +1 | 2e-116 | 353 | 187/354 (53%) | DNA-binding protein phosphatase 1 | chr2:10903154-10904978 REVERSE LENGTH=392 |
