Probe CUST_20770_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20770_PI426222305 | JHI_St_60k_v1 | DMT400011761 | CTCCTCAAGCTTGGAAAGATCCTTTGAAGAATTGTGCCTTTTCATATAAGGTAATGTTAT |
All Microarray Probes Designed to Gene DMG400004617
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20924_PI426222305 | JHI_St_60k_v1 | DMT400011762 | TGTCCCCTTCTAGATAAGTCTTTACTATAACCACTGAAAACACAAAGCAAGAGATGATTA |
| CUST_20770_PI426222305 | JHI_St_60k_v1 | DMT400011761 | CTCCTCAAGCTTGGAAAGATCCTTTGAAGAATTGTGCCTTTTCATATAAGGTAATGTTAT |
Microarray Signals from CUST_20770_PI426222305
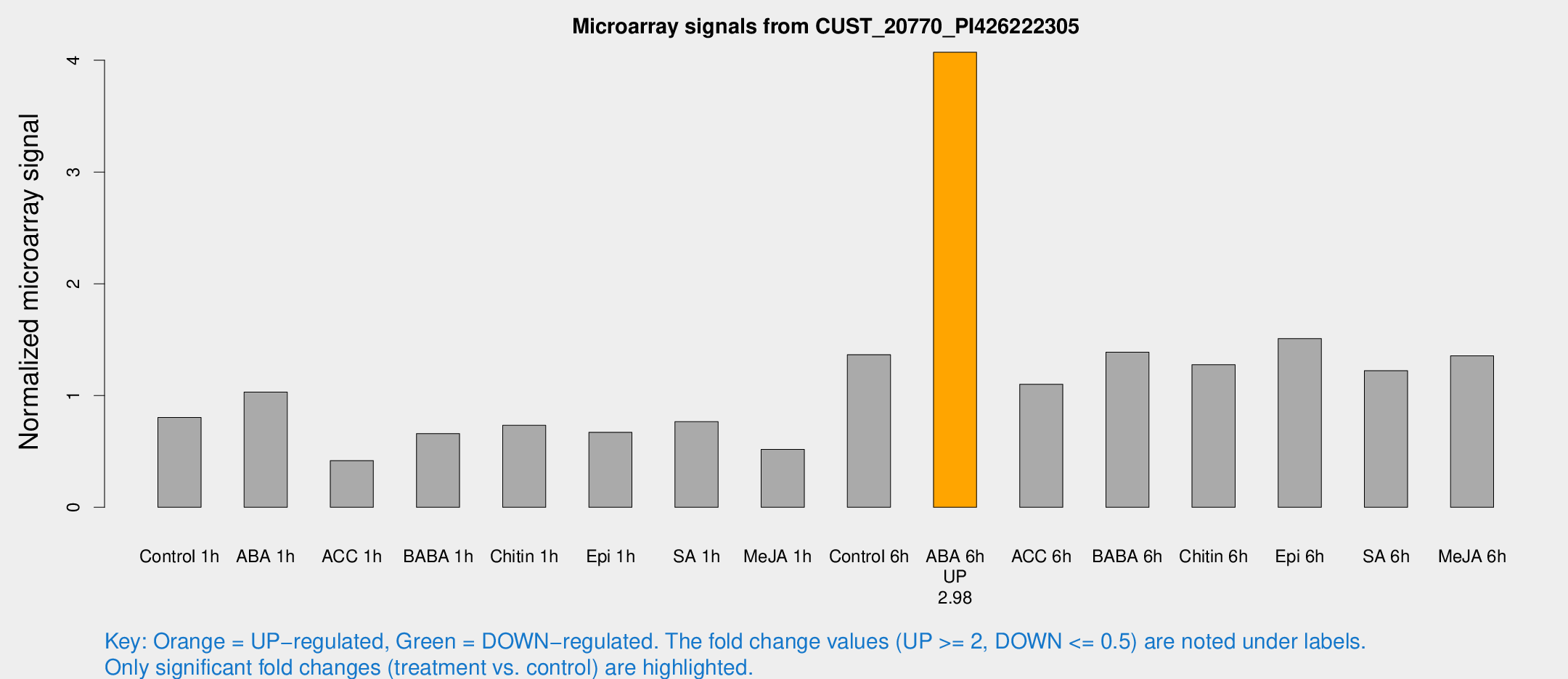
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1309.4 | 97.481 | 0.803667 | 0.0929737 |
| ABA 1h | 1532.73 | 274.47 | 1.03011 | 0.14907 |
| ACC 1h | 752.056 | 189.409 | 0.417291 | 0.0966586 |
| BABA 1h | 1116.84 | 282.591 | 0.660064 | 0.132035 |
| Chitin 1h | 1105.63 | 192.474 | 0.733943 | 0.136693 |
| Epi 1h | 975.873 | 172.301 | 0.670968 | 0.127248 |
| SA 1h | 1294.68 | 161.569 | 0.766268 | 0.075971 |
| Me-JA 1h | 689.166 | 55.4054 | 0.517742 | 0.0299915 |
| Control 6h | 2324.01 | 488.878 | 1.36545 | 0.207596 |
| ABA 6h | 7179.21 | 1043.2 | 4.0718 | 0.440677 |
| ACC 6h | 2100.08 | 324.105 | 1.10113 | 0.102325 |
| BABA 6h | 2522.9 | 146.128 | 1.38717 | 0.0801156 |
| Chitin 6h | 2202.31 | 127.313 | 1.27606 | 0.0737073 |
| Epi 6h | 2778.95 | 247.623 | 1.5096 | 0.0871822 |
| SA 6h | 2078.89 | 453.863 | 1.22227 | 0.169261 |
| Me-JA 6h | 2316.96 | 527.964 | 1.35472 | 0.225929 |
Source Transcript PGSC0003DMT400011761 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G47960.1 | +3 | 2e-11 | 59 | 33/81 (41%) | cell wall / vacuolar inhibitor of fructosidase 1 | chr1:17681954-17683516 REVERSE LENGTH=205 |
