Probe CUST_20742_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20742_PI426222305 | JHI_St_60k_v1 | DMT400011902 | TCTATGGACAAACTCTCTGGCTCTGGTTTTCAATCCAAAAAAGATTATCTTTTTGGGAGA |
All Microarray Probes Designed to Gene DMG400004670
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20742_PI426222305 | JHI_St_60k_v1 | DMT400011902 | TCTATGGACAAACTCTCTGGCTCTGGTTTTCAATCCAAAAAAGATTATCTTTTTGGGAGA |
| CUST_20748_PI426222305 | JHI_St_60k_v1 | DMT400011901 | GCAGCTATCTTCTAAAGGATCAACTCATGATGAAATTGACTTTGAGTTCTTAGGAAATGT |
Microarray Signals from CUST_20742_PI426222305
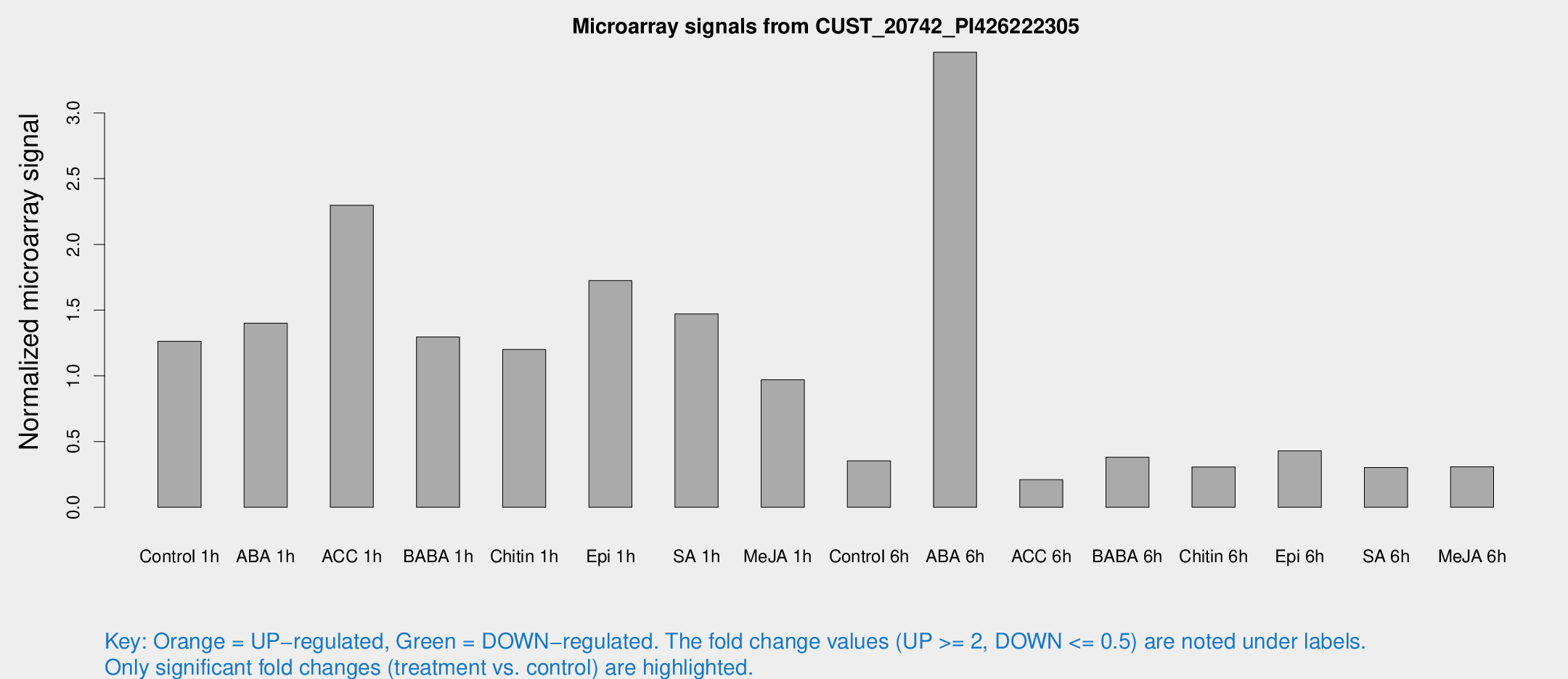
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 86.1414 | 23.6196 | 1.26312 | 0.292523 |
| ABA 1h | 79.2641 | 10.6599 | 1.40025 | 0.265743 |
| ACC 1h | 170.066 | 55.5201 | 2.29714 | 0.775971 |
| BABA 1h | 79.6821 | 9.41587 | 1.29611 | 0.293086 |
| Chitin 1h | 68.1731 | 5.28905 | 1.2001 | 0.146003 |
| Epi 1h | 94.8742 | 9.35816 | 1.72456 | 0.159949 |
| SA 1h | 95.8165 | 8.60254 | 1.47049 | 0.135213 |
| Me-JA 1h | 51.7142 | 9.13622 | 0.970692 | 0.283727 |
| Control 6h | 31.4869 | 15.6299 | 0.353995 | 0.246486 |
| ABA 6h | 286.121 | 120.659 | 3.4619 | 2.14343 |
| ACC 6h | 19.9394 | 10.7042 | 0.21006 | 0.0861839 |
| BABA 6h | 26.9415 | 4.43306 | 0.381593 | 0.0623947 |
| Chitin 6h | 20.9372 | 4.28389 | 0.307241 | 0.0666948 |
| Epi 6h | 41.4862 | 18.9226 | 0.428841 | 0.412786 |
| SA 6h | 23.2561 | 8.29155 | 0.302416 | 0.237603 |
| Me-JA 6h | 20.0806 | 3.92981 | 0.308027 | 0.0751296 |
Source Transcript PGSC0003DMT400011902 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14130.1 | +2 | 5e-146 | 419 | 198/265 (75%) | xyloglucan endotransglucosylase/hydrolase 15 | chr4:8137161-8138196 REVERSE LENGTH=289 |
