Probe CUST_20378_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20378_PI426222305 | JHI_St_60k_v1 | DMT400049802 | CCCAGATTCTTCAAACATAATAATTTCTCGAGCTTTGTCAGACAGCAAAATACGTATGTA |
All Microarray Probes Designed to Gene DMG400019344
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20297_PI426222305 | JHI_St_60k_v1 | DMT400049803 | GTATCTTTTGACTTAATTTTGTGTGAAGAAGAGTGTATGATCAGTGATTGCTGACTCTTC |
| CUST_20378_PI426222305 | JHI_St_60k_v1 | DMT400049802 | CCCAGATTCTTCAAACATAATAATTTCTCGAGCTTTGTCAGACAGCAAAATACGTATGTA |
Microarray Signals from CUST_20378_PI426222305
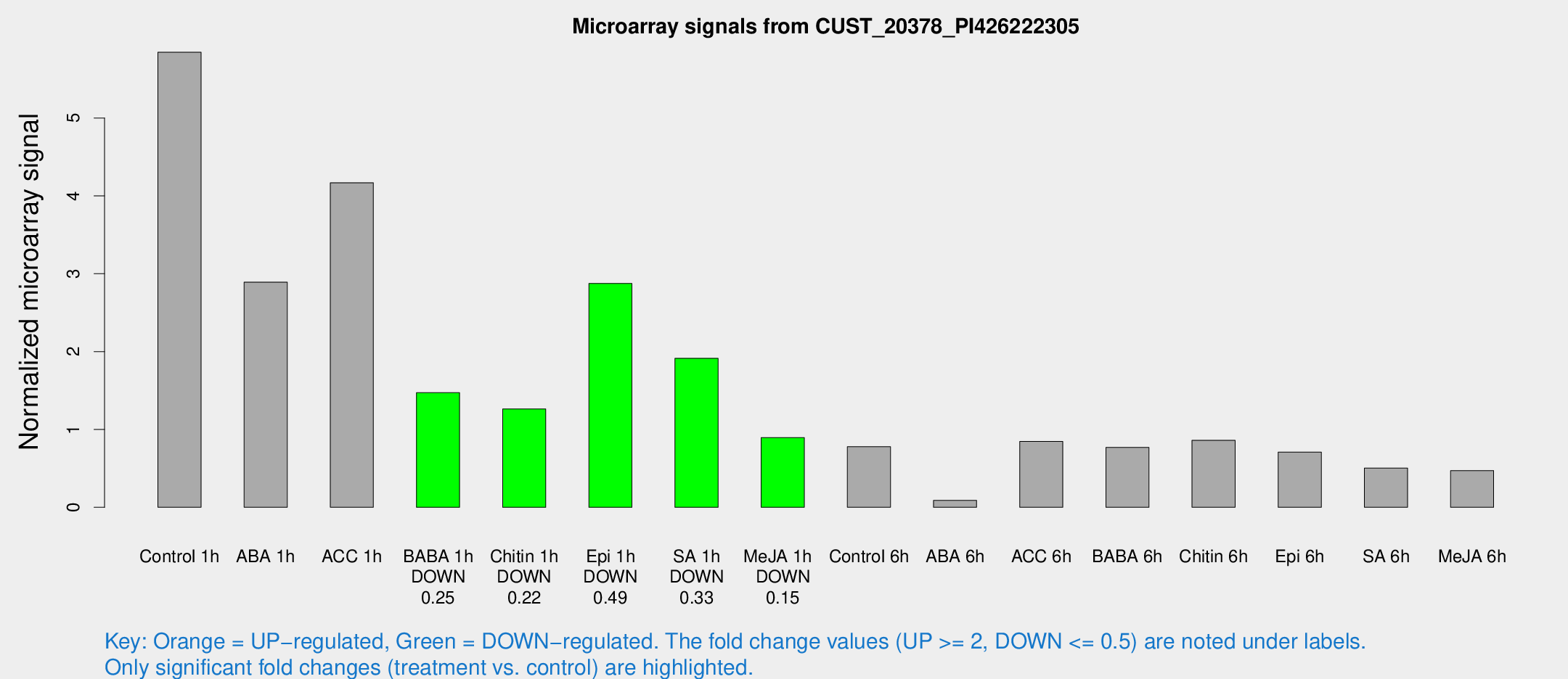
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1477.61 | 187.756 | 5.84483 | 0.352357 |
| ABA 1h | 687.201 | 194.699 | 2.89219 | 0.939395 |
| ACC 1h | 1150.18 | 306.59 | 4.16789 | 0.946055 |
| BABA 1h | 407.521 | 132.972 | 1.4726 | 0.476193 |
| Chitin 1h | 283.567 | 16.7631 | 1.26254 | 0.125586 |
| Epi 1h | 627.299 | 68.0229 | 2.87458 | 0.410868 |
| SA 1h | 498.686 | 70.6989 | 1.91346 | 0.154389 |
| Me-JA 1h | 185.533 | 24.6808 | 0.895902 | 0.054361 |
| Control 6h | 208.468 | 51.5362 | 0.777995 | 0.156064 |
| ABA 6h | 26.1503 | 7.4741 | 0.0892019 | 0.0270652 |
| ACC 6h | 245.24 | 31.362 | 0.845121 | 0.0507725 |
| BABA 6h | 234.973 | 61.8408 | 0.768307 | 0.236237 |
| Chitin 6h | 232.685 | 32.3704 | 0.860446 | 0.116322 |
| Epi 6h | 211.601 | 54.4785 | 0.709255 | 0.192143 |
| SA 6h | 127.404 | 20.6642 | 0.504234 | 0.146968 |
| Me-JA 6h | 118.073 | 12.7597 | 0.471058 | 0.0598667 |
Source Transcript PGSC0003DMT400049802 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G51910.1 | +1 | 2e-35 | 122 | 54/70 (77%) | heat shock transcription factor A7A | chr3:19265294-19266619 FORWARD LENGTH=272 |
