Probe CUST_20349_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20349_PI426222305 | JHI_St_60k_v1 | DMT400049791 | CTTCCCAGATTCTTCAAACACAACAATTTCTCAAGCTTTGTCAGACAGCTAAATACTTAT |
All Microarray Probes Designed to Gene DMG402019343
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20349_PI426222305 | JHI_St_60k_v1 | DMT400049791 | CTTCCCAGATTCTTCAAACACAACAATTTCTCAAGCTTTGTCAGACAGCTAAATACTTAT |
| CUST_20361_PI426222305 | JHI_St_60k_v1 | DMT400049793 | CTTCCCAGATTCTTCAAACACAACAATTTCTCAAGCTTTGTCAGACAGCTAAATACTTAT |
Microarray Signals from CUST_20349_PI426222305
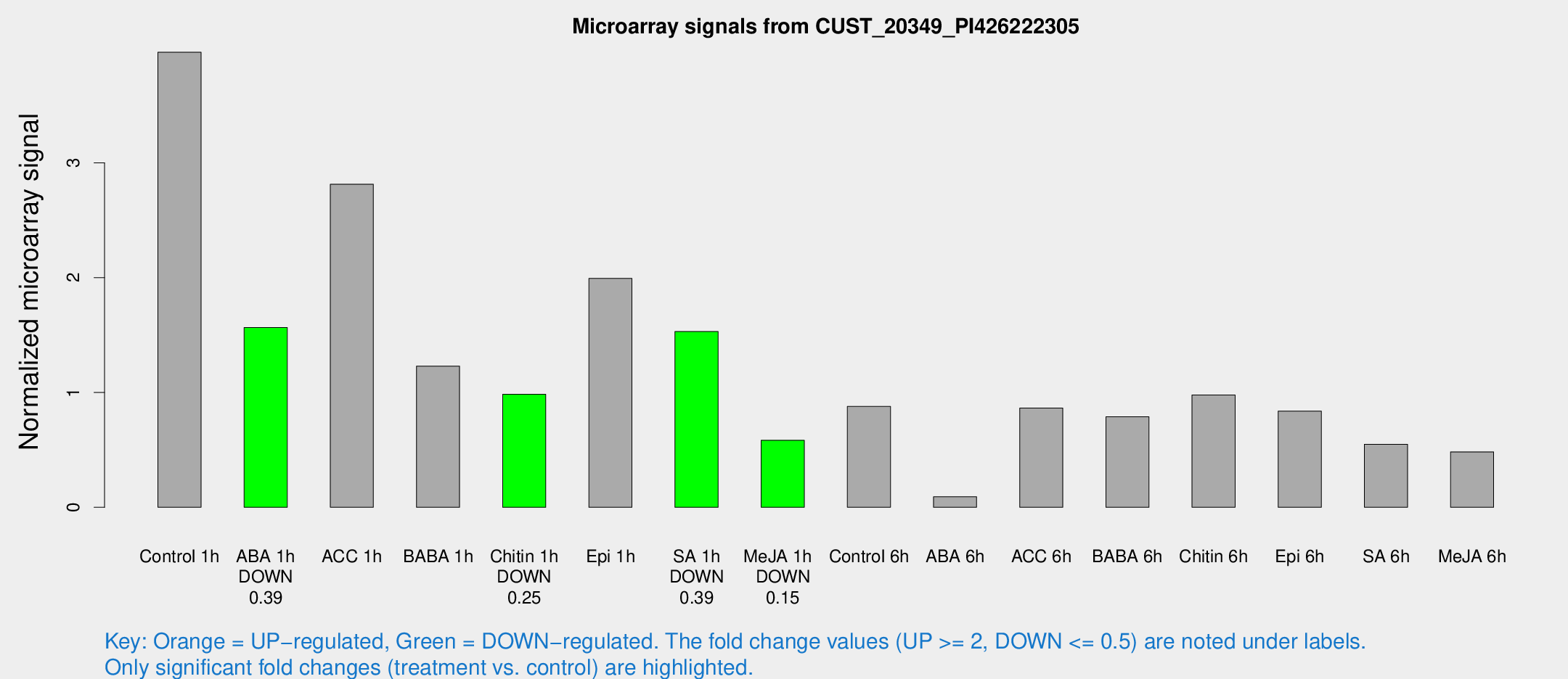
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1258.36 | 214.679 | 3.96269 | 0.459348 |
| ABA 1h | 432.94 | 48.6393 | 1.56471 | 0.198921 |
| ACC 1h | 944.057 | 208.55 | 2.81336 | 0.499586 |
| BABA 1h | 432.516 | 145.111 | 1.22917 | 0.458115 |
| Chitin 1h | 273.701 | 16.2338 | 0.984033 | 0.104986 |
| Epi 1h | 533.188 | 30.9981 | 1.99285 | 0.11585 |
| SA 1h | 509.942 | 112.848 | 1.53065 | 0.298781 |
| Me-JA 1h | 151.365 | 25.7708 | 0.583792 | 0.0661002 |
| Control 6h | 286.132 | 63.2486 | 0.87856 | 0.14455 |
| ABA 6h | 35.8217 | 12.86 | 0.0919449 | 0.0402148 |
| ACC 6h | 316.947 | 60.2485 | 0.864541 | 0.051513 |
| BABA 6h | 295.868 | 73.763 | 0.788062 | 0.221434 |
| Chitin 6h | 337.508 | 67.2896 | 0.978748 | 0.22427 |
| Epi 6h | 316.024 | 93.1095 | 0.838059 | 0.234681 |
| SA 6h | 171.163 | 22.9924 | 0.549193 | 0.141472 |
| Me-JA 6h | 148.745 | 9.39989 | 0.482846 | 0.0309072 |
Source Transcript PGSC0003DMT400049791 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G22830.1 | +1 | 7e-66 | 225 | 130/280 (46%) | heat shock transcription factor A6B | chr3:8078981-8080895 FORWARD LENGTH=406 |
