Probe CUST_19609_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19609_PI426222305 | JHI_St_60k_v1 | DMT400023251 | AGATTGTATGCTCAGATGTCTTATTTGGGAAGTTTGGCTTATGGAATACCCCAAATTAAG |
All Microarray Probes Designed to Gene DMG400009005
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19588_PI426222305 | JHI_St_60k_v1 | DMT400023252 | CTGGTTTTCTCAAAAGGGAACTTCCTGGATGAATGAAATCCAAATAGATTTACAATTTGG |
| CUST_19609_PI426222305 | JHI_St_60k_v1 | DMT400023251 | AGATTGTATGCTCAGATGTCTTATTTGGGAAGTTTGGCTTATGGAATACCCCAAATTAAG |
Microarray Signals from CUST_19609_PI426222305
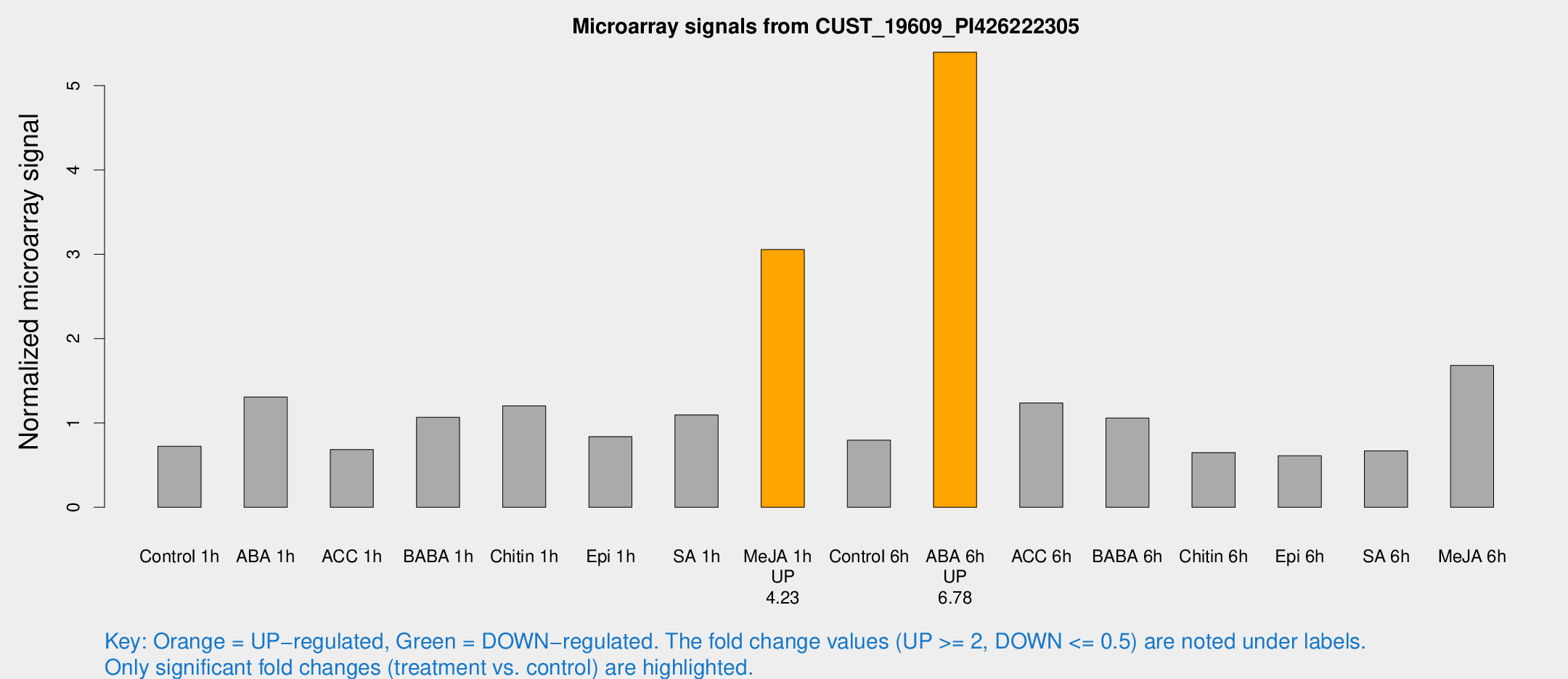
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 84.5807 | 5.90765 | 0.722588 | 0.0504902 |
| ABA 1h | 143.348 | 33.1034 | 1.30891 | 0.292402 |
| ACC 1h | 93.6868 | 32.2894 | 0.684739 | 0.227812 |
| BABA 1h | 127.521 | 32.2614 | 1.06637 | 0.193965 |
| Chitin 1h | 127.384 | 12.4996 | 1.20201 | 0.150453 |
| Epi 1h | 84.7846 | 5.85714 | 0.837418 | 0.0578843 |
| SA 1h | 150.915 | 58.2946 | 1.09581 | 0.424443 |
| Me-JA 1h | 296.696 | 40.2732 | 3.05539 | 0.179904 |
| Control 6h | 104.525 | 34.1799 | 0.796201 | 0.21443 |
| ABA 6h | 784.296 | 265.54 | 5.39608 | 2.26722 |
| ACC 6h | 166.134 | 10.4266 | 1.23648 | 0.104597 |
| BABA 6h | 159.874 | 63.2912 | 1.05857 | 0.386037 |
| Chitin 6h | 80.9157 | 6.00182 | 0.648936 | 0.0545281 |
| Epi 6h | 109.396 | 53.1092 | 0.611404 | 0.54282 |
| SA 6h | 111.367 | 46.3591 | 0.670331 | 0.883717 |
| Me-JA 6h | 198.544 | 23.6827 | 1.68131 | 0.101559 |
Source Transcript PGSC0003DMT400023251 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G02660.1 | +3 | 3e-178 | 536 | 359/581 (62%) | alpha/beta-Hydrolases superfamily protein | chr1:572187-574746 REVERSE LENGTH=713 |
