Probe CUST_18819_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18819_PI426222305 | JHI_St_60k_v1 | DMT400001076 | GATGGTCCCATTTGCCATTAAAAGTTGCAAAAAGTAGTTGAAGCGTCCAAGGTGTTTTTT |
All Microarray Probes Designed to Gene DMG400000408
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18819_PI426222305 | JHI_St_60k_v1 | DMT400001076 | GATGGTCCCATTTGCCATTAAAAGTTGCAAAAAGTAGTTGAAGCGTCCAAGGTGTTTTTT |
Microarray Signals from CUST_18819_PI426222305
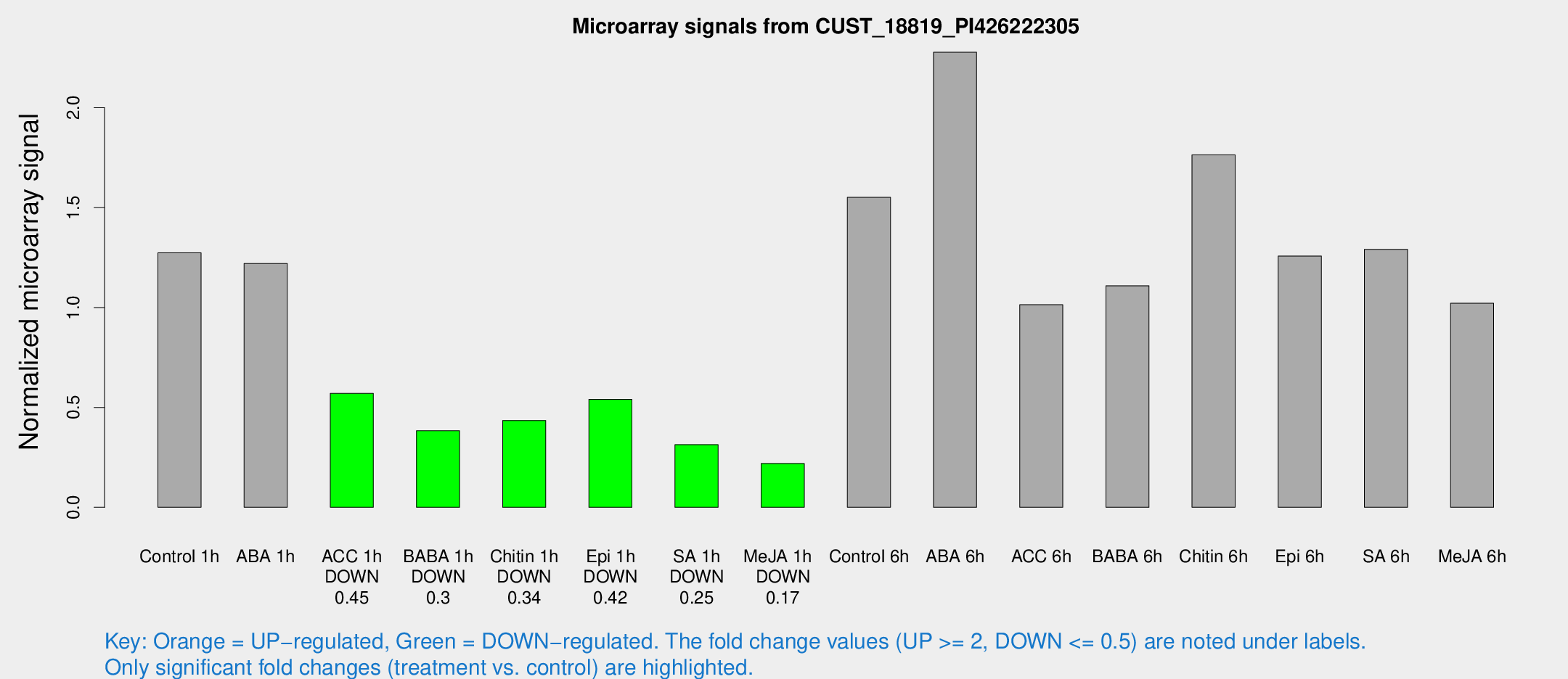
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5396.72 | 312.61 | 1.27368 | 0.07354 |
| ABA 1h | 4665.41 | 736.98 | 1.22024 | 0.218894 |
| ACC 1h | 2577.3 | 483.881 | 0.570068 | 0.0784206 |
| BABA 1h | 1595.98 | 235.867 | 0.383283 | 0.0267746 |
| Chitin 1h | 1720.72 | 368.152 | 0.434567 | 0.0619806 |
| Epi 1h | 2054.93 | 390.584 | 0.540377 | 0.109117 |
| SA 1h | 1386.18 | 188.556 | 0.313767 | 0.0382976 |
| Me-JA 1h | 761.436 | 59.7883 | 0.219478 | 0.0171535 |
| Control 6h | 7138.19 | 2168.72 | 1.5519 | 0.422702 |
| ABA 6h | 10294.8 | 804.278 | 2.27827 | 0.131538 |
| ACC 6h | 5034.53 | 741.263 | 1.01402 | 0.148488 |
| BABA 6h | 5798.95 | 1624.72 | 1.10922 | 0.444362 |
| Chitin 6h | 7973.82 | 556.303 | 1.76483 | 0.14941 |
| Epi 6h | 6026.2 | 458.696 | 1.25817 | 0.112818 |
| SA 6h | 5603.06 | 1002.19 | 1.29151 | 0.106747 |
| Me-JA 6h | 4716.05 | 1511.22 | 1.02147 | 0.259064 |
Source Transcript PGSC0003DMT400001076 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36870.1 | +3 | 4e-128 | 376 | 197/279 (71%) | xyloglucan endotransglucosylase/hydrolase 32 | chr2:15472869-15474630 REVERSE LENGTH=299 |
