Probe CUST_18745_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18745_PI426222305 | JHI_St_60k_v1 | DMT400001154 | GGTATTCATGTTTTTGTTTCATAAGTATACTAGCTGTAAAATGGAGGTCTAGTGGGACTA |
All Microarray Probes Designed to Gene DMG400000434
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18876_PI426222305 | JHI_St_60k_v1 | DMT400001153 | GGTATTCATGTTTTTGTTTCATAAGTATACTAGCTGTAAAATGGAGGTCTAGTGGGACTA |
| CUST_18745_PI426222305 | JHI_St_60k_v1 | DMT400001154 | GGTATTCATGTTTTTGTTTCATAAGTATACTAGCTGTAAAATGGAGGTCTAGTGGGACTA |
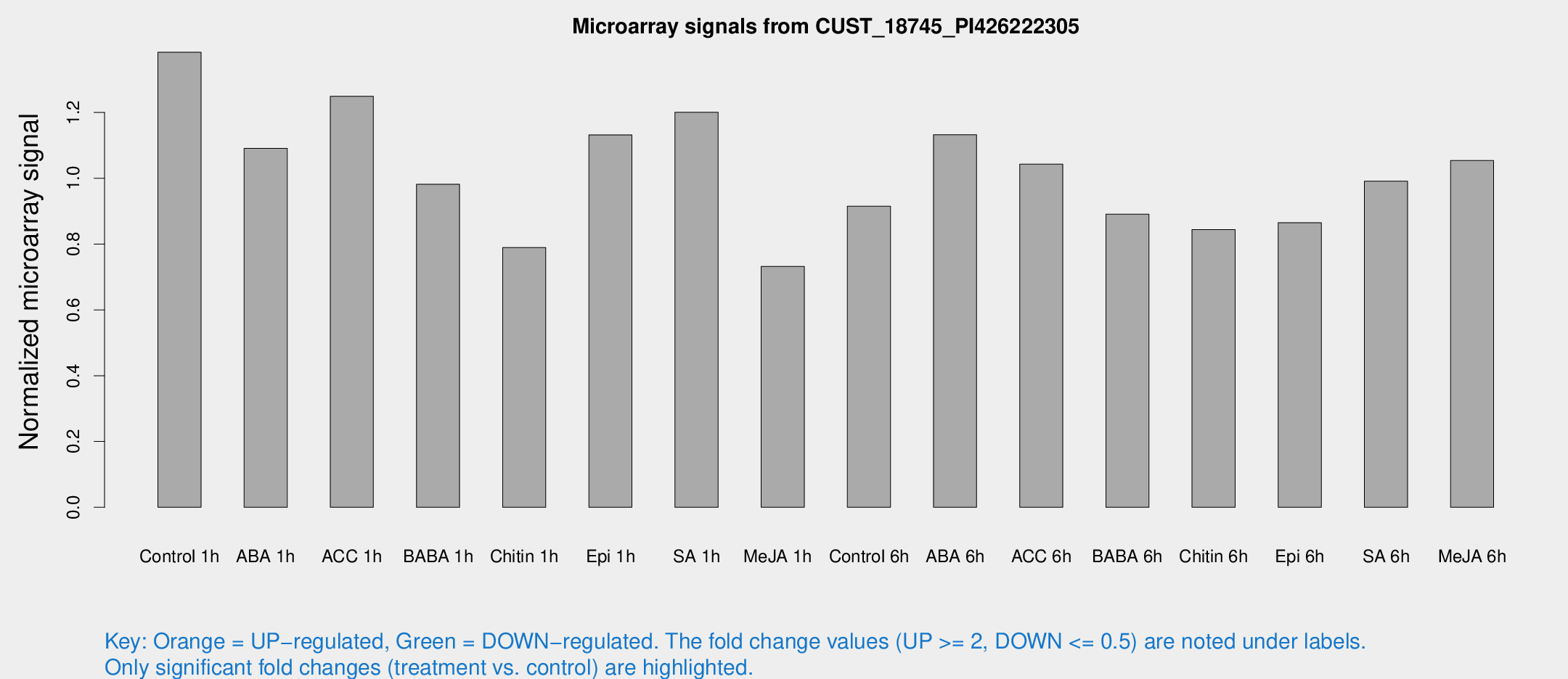
Microarray Signals from CUST_18745_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 307.545 | 51.8235 | 1.38307 | 0.13743 |
| ABA 1h | 209.306 | 12.4635 | 1.09105 | 0.119344 |
| ACC 1h | 279.509 | 21.4153 | 1.24923 | 0.0743277 |
| BABA 1h | 208.3 | 26.0199 | 0.981665 | 0.0589596 |
| Chitin 1h | 154.63 | 11.7012 | 0.789574 | 0.107818 |
| Epi 1h | 212.819 | 12.6932 | 1.13177 | 0.0673465 |
| SA 1h | 267.346 | 15.7675 | 1.20057 | 0.0784977 |
| Me-JA 1h | 130.841 | 13.4011 | 0.732362 | 0.0694923 |
| Control 6h | 204.168 | 32.6039 | 0.914933 | 0.0861659 |
| ABA 6h | 263.707 | 27.8232 | 1.13248 | 0.0669602 |
| ACC 6h | 260.595 | 20.3481 | 1.04287 | 0.196759 |
| BABA 6h | 217.807 | 20.382 | 0.890623 | 0.0534978 |
| Chitin 6h | 194.925 | 11.8161 | 0.843929 | 0.051095 |
| Epi 6h | 211.602 | 12.8002 | 0.864953 | 0.0882423 |
| SA 6h | 220.876 | 43.8224 | 0.990944 | 0.112705 |
| Me-JA 6h | 229.895 | 23.8189 | 1.05376 | 0.0626581 |
Source Transcript PGSC0003DMT400001154 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G44820.1 | +2 | 0.0 | 718 | 395/632 (63%) | Phototropic-responsive NPH3 family protein | chr3:16361864-16364411 REVERSE LENGTH=651 |
