Probe CUST_18468_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18468_PI426222305 | JHI_St_60k_v1 | DMT400005248 | GATGTTAAGAAATATGGTTATAAAGTAGGTCTCCCTAGGTTTCTTGTCTCAACTCACTAA |
All Microarray Probes Designed to Gene DMG400002068
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18468_PI426222305 | JHI_St_60k_v1 | DMT400005248 | GATGTTAAGAAATATGGTTATAAAGTAGGTCTCCCTAGGTTTCTTGTCTCAACTCACTAA |
| CUST_18453_PI426222305 | JHI_St_60k_v1 | DMT400005250 | GGTCTCCCTAGGTTTCTTGTCTCAACTCACTAAACTCATATAAAATTTTGTAGCAGTTAA |
Microarray Signals from CUST_18468_PI426222305
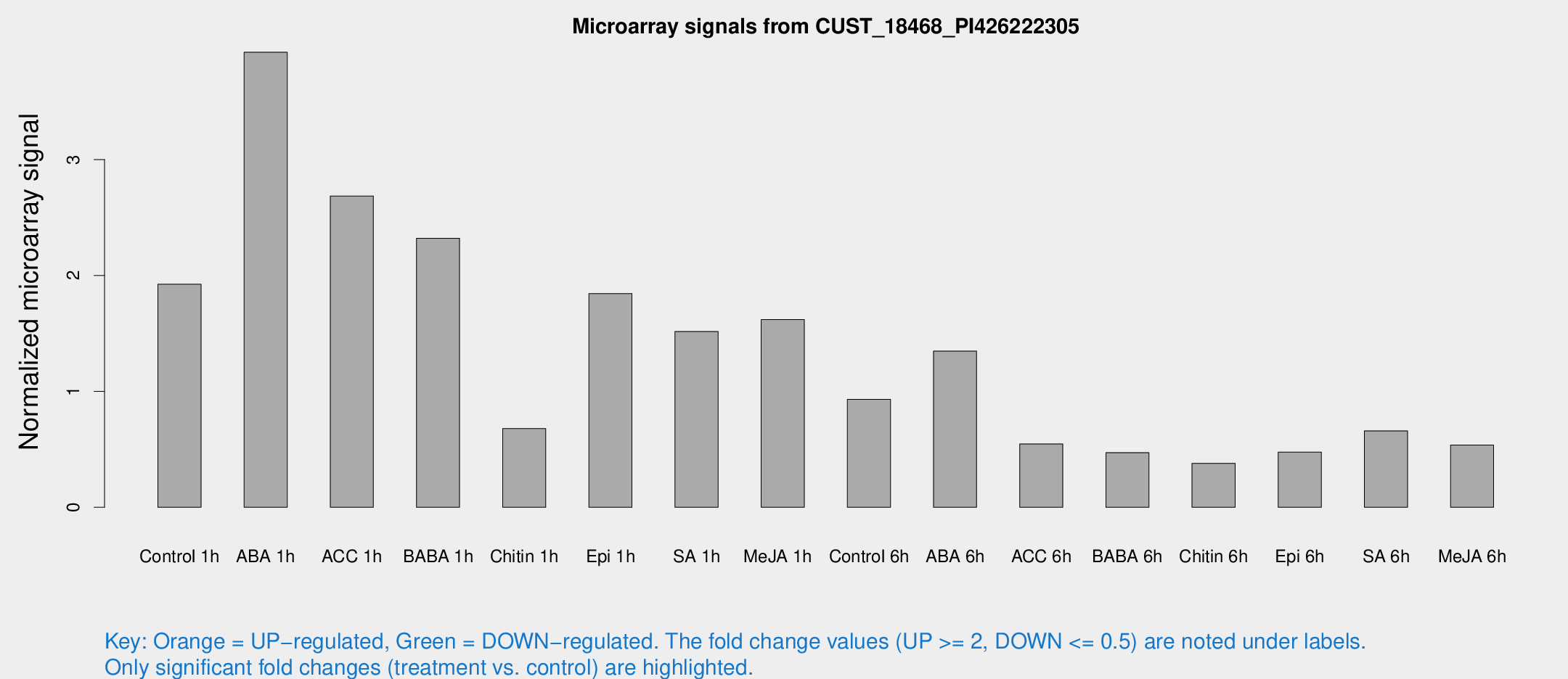
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 708.663 | 112.242 | 1.92446 | 0.385334 |
| ABA 1h | 1286.99 | 230.206 | 3.92554 | 0.491035 |
| ACC 1h | 1044.02 | 222.528 | 2.68457 | 0.466515 |
| BABA 1h | 818.862 | 125.201 | 2.32122 | 0.613641 |
| Chitin 1h | 228.584 | 47.3334 | 0.67923 | 0.128042 |
| Epi 1h | 585.235 | 94.3257 | 1.84406 | 0.301753 |
| SA 1h | 574.376 | 93.8519 | 1.51648 | 0.375168 |
| Me-JA 1h | 478.582 | 50.1767 | 1.61894 | 0.191549 |
| Control 6h | 383.65 | 135.917 | 0.930868 | 0.309706 |
| ABA 6h | 542.727 | 131.319 | 1.34777 | 0.314583 |
| ACC 6h | 271.249 | 122.635 | 0.546506 | 0.171425 |
| BABA 6h | 194.49 | 33.699 | 0.471706 | 0.0849504 |
| Chitin 6h | 144.645 | 8.97481 | 0.379233 | 0.0235157 |
| Epi 6h | 197.511 | 30.4349 | 0.47644 | 0.113721 |
| SA 6h | 236.457 | 25.7379 | 0.659745 | 0.0391713 |
| Me-JA 6h | 192.673 | 15.5748 | 0.536972 | 0.0787529 |
Source Transcript PGSC0003DMT400005248 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G21200.1 | +1 | 3e-146 | 419 | 201/302 (67%) | gibberellin 2-oxidase 8 | chr4:11302751-11306601 FORWARD LENGTH=338 |
