Probe CUST_18420_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18420_PI426222305 | JHI_St_60k_v1 | DMT400042447 | GAAGTGACTCTTTTTTGGACACCCTTTGCCTTTTTCTGTATTGGGTATTAATTTTCTTGA |
All Microarray Probes Designed to Gene DMG400016460
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18621_PI426222305 | JHI_St_60k_v1 | DMT400042448 | CTGAGAATTATGTTCACTGGAGAGAGACCCCTGAATCTCACATATACTCTGCTGATCTTC |
| CUST_18420_PI426222305 | JHI_St_60k_v1 | DMT400042447 | GAAGTGACTCTTTTTTGGACACCCTTTGCCTTTTTCTGTATTGGGTATTAATTTTCTTGA |
Microarray Signals from CUST_18420_PI426222305
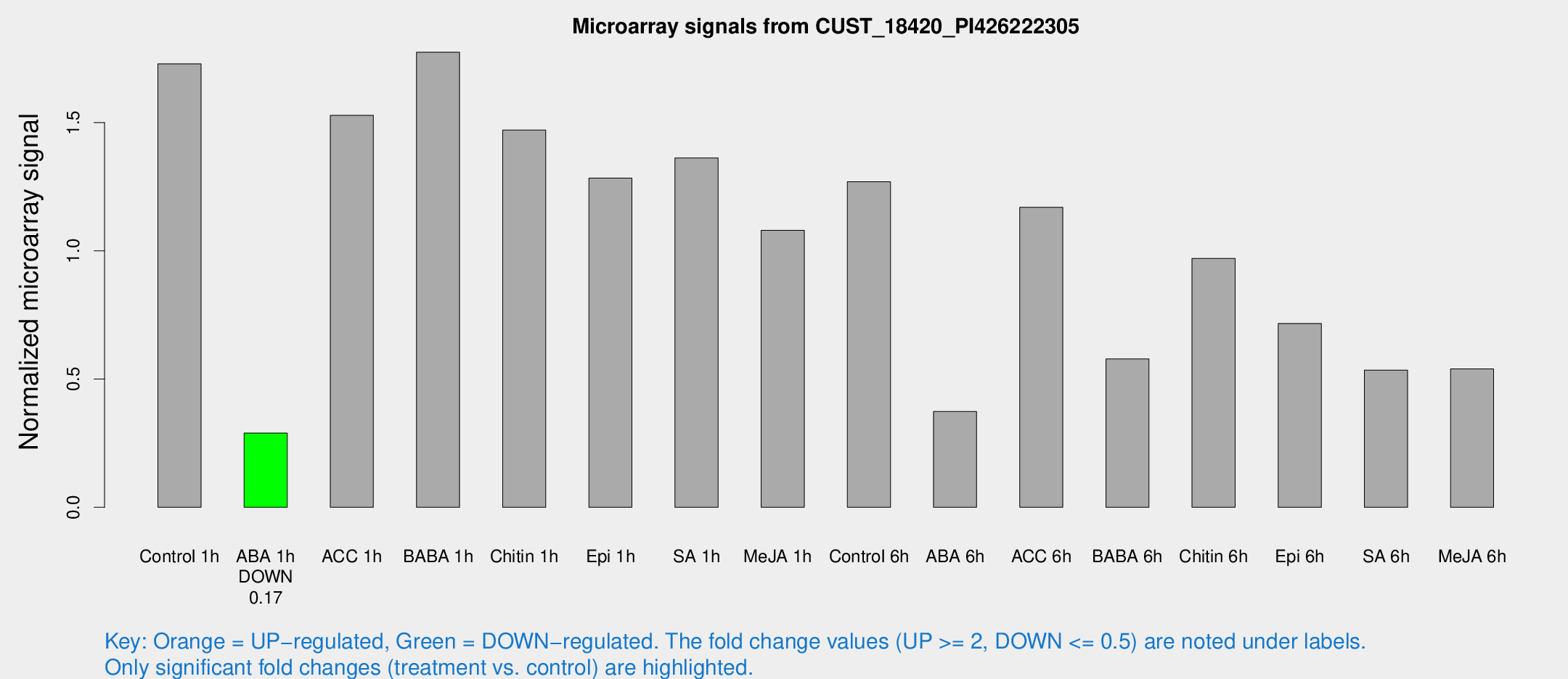
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 37.4559 | 6.24311 | 1.72912 | 0.274655 |
| ABA 1h | 5.38441 | 3.02801 | 0.289201 | 0.162711 |
| ACC 1h | 39.4872 | 13.3334 | 1.52831 | 0.666693 |
| BABA 1h | 42.5335 | 13.9652 | 1.77461 | 0.669342 |
| Chitin 1h | 31.2384 | 9.62297 | 1.47113 | 0.696349 |
| Epi 1h | 24.8112 | 5.42551 | 1.28359 | 0.325208 |
| SA 1h | 30.0692 | 4.28746 | 1.36217 | 0.17254 |
| Me-JA 1h | 21.4211 | 6.75527 | 1.07995 | 0.507347 |
| Control 6h | 29.7057 | 8.15363 | 1.26992 | 0.334217 |
| ABA 6h | 9.33442 | 3.39181 | 0.373798 | 0.16615 |
| ACC 6h | 30.9428 | 9.84372 | 1.17023 | 0.189787 |
| BABA 6h | 14.5264 | 3.72645 | 0.578502 | 0.17103 |
| Chitin 6h | 22.1693 | 3.81778 | 0.970761 | 0.176576 |
| Epi 6h | 18.7207 | 5.65889 | 0.71688 | 0.277393 |
| SA 6h | 12.0864 | 3.53827 | 0.53518 | 0.202347 |
| Me-JA 6h | 14.3686 | 7.18013 | 0.539514 | 0.289478 |
Source Transcript PGSC0003DMT400042447 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G21870.1 | +1 | 4e-15 | 72 | 70/183 (38%) | HSP20-like chaperones superfamily protein | chr4:11603756-11604285 REVERSE LENGTH=134 |
