Probe CUST_18167_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18167_PI426222305 | JHI_St_60k_v1 | DMT400048453 | GGATGTCTTCAAGATACTAGTCTTGAATCTTGATCTTCTTAGAAATGTGAAGCAGTGATT |
All Microarray Probes Designed to Gene DMG400018819
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18141_PI426222305 | JHI_St_60k_v1 | DMT400048452 | CTTGGCTTCTCCTTTTTTAATTGGAGTAAGTCGAAATGGTTCTCTGATATTAGTAAGATG |
| CUST_18167_PI426222305 | JHI_St_60k_v1 | DMT400048453 | GGATGTCTTCAAGATACTAGTCTTGAATCTTGATCTTCTTAGAAATGTGAAGCAGTGATT |
Microarray Signals from CUST_18167_PI426222305
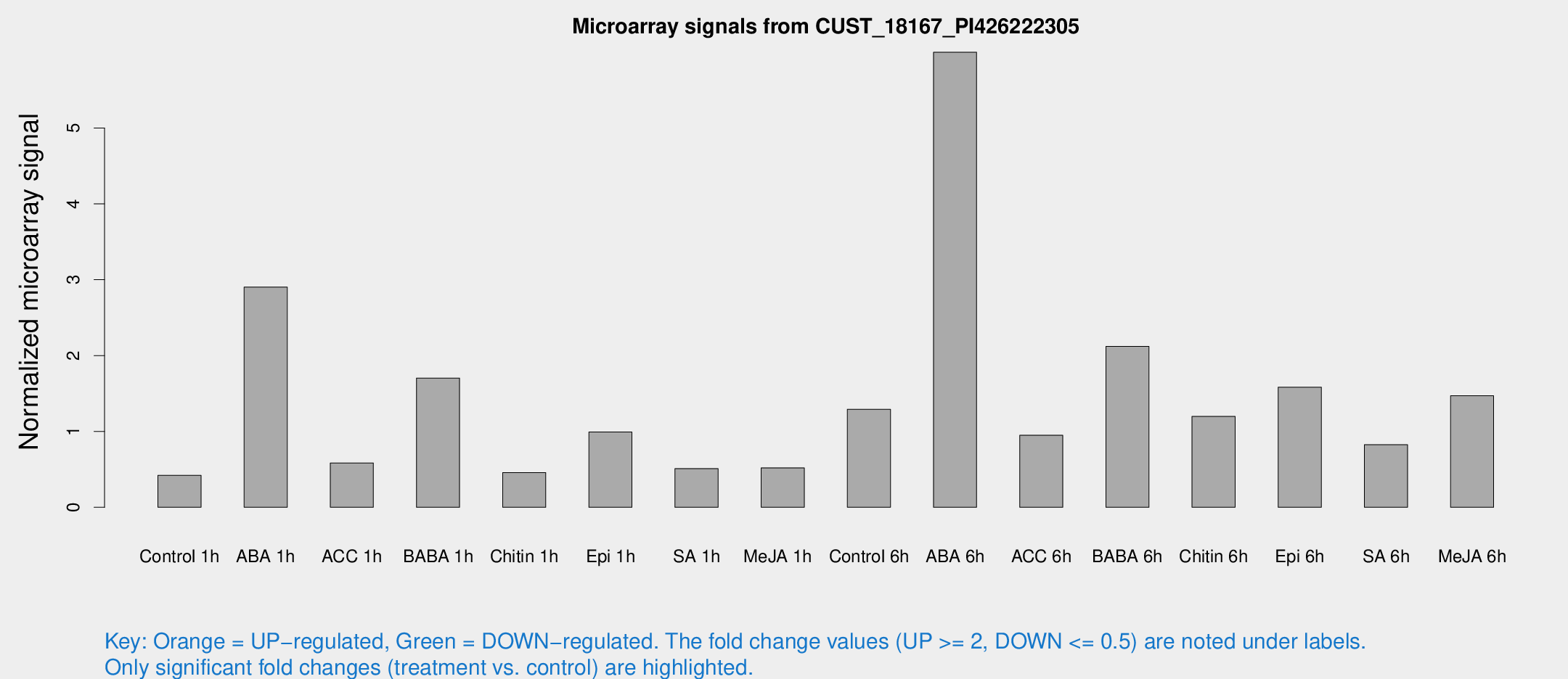
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.85456 | 3.35699 | 0.422382 | 0.242128 |
| ABA 1h | 41.6867 | 13.7761 | 2.90364 | 1.26094 |
| ACC 1h | 8.33824 | 4.08823 | 0.584519 | 0.282424 |
| BABA 1h | 24.4656 | 6.00949 | 1.70403 | 0.32016 |
| Chitin 1h | 5.71518 | 3.31825 | 0.458421 | 0.265552 |
| Epi 1h | 13.3509 | 4.14238 | 0.992268 | 0.364505 |
| SA 1h | 7.67587 | 3.35088 | 0.510596 | 0.250025 |
| Me-JA 1h | 5.88932 | 3.42487 | 0.52025 | 0.301679 |
| Control 6h | 24.678 | 12.4002 | 1.29275 | 0.845874 |
| ABA 6h | 100.954 | 31.643 | 5.99903 | 2.36566 |
| ACC 6h | 17.741 | 5.77138 | 0.950141 | 0.397556 |
| BABA 6h | 39.8313 | 15.7753 | 2.12107 | 1.03202 |
| Chitin 6h | 19.0597 | 5.47347 | 1.19844 | 0.313394 |
| Epi 6h | 32.3973 | 13.5329 | 1.58345 | 1.39342 |
| SA 6h | 12.1231 | 3.67253 | 0.825459 | 0.304967 |
| Me-JA 6h | 25.6877 | 10.096 | 1.4703 | 0.792185 |
Source Transcript PGSC0003DMT400048453 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G08570.1 | +1 | 3e-71 | 227 | 106/146 (73%) | atypical CYS HIS rich thioredoxin 4 | chr1:2713059-2714312 FORWARD LENGTH=275 |
