Probe CUST_18027_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18027_PI426222305 | JHI_St_60k_v1 | DMT400071105 | GGAGTTGCCTACTTTAGATGATCTTCTTCGTCTTTATTCAGACGAAATACAACAATAACA |
All Microarray Probes Designed to Gene DMG400027649
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18027_PI426222305 | JHI_St_60k_v1 | DMT400071105 | GGAGTTGCCTACTTTAGATGATCTTCTTCGTCTTTATTCAGACGAAATACAACAATAACA |
Microarray Signals from CUST_18027_PI426222305
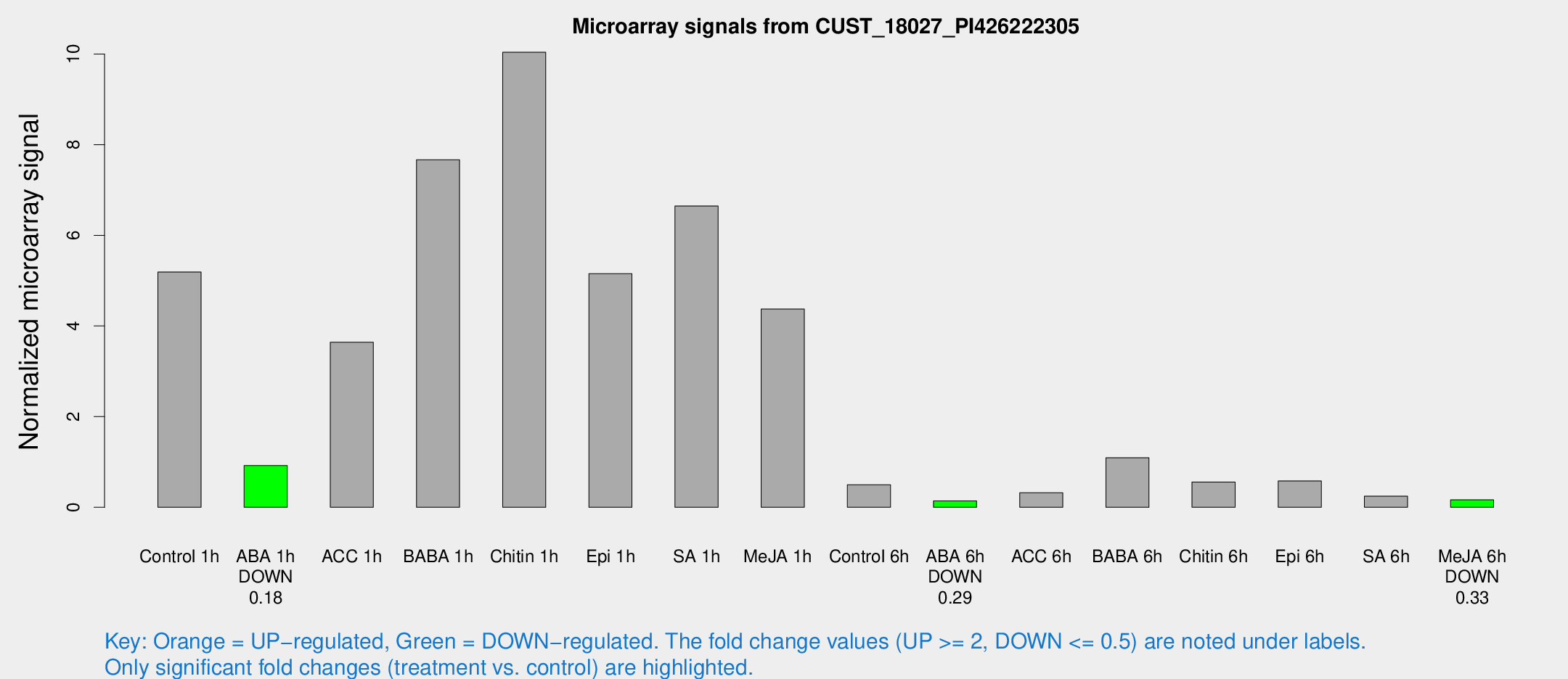
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 431.233 | 73.7859 | 5.19042 | 0.840488 |
| ABA 1h | 66.1724 | 6.1807 | 0.920134 | 0.0805255 |
| ACC 1h | 392.699 | 150.23 | 3.642 | 2.27619 |
| BABA 1h | 610.019 | 89.4725 | 7.66528 | 0.444465 |
| Chitin 1h | 737.853 | 90.9704 | 10.0391 | 1.98355 |
| Epi 1h | 363.701 | 39.294 | 5.15307 | 0.481668 |
| SA 1h | 563.134 | 80.891 | 6.64739 | 1.20943 |
| Me-JA 1h | 298.073 | 53.8837 | 4.37362 | 0.806293 |
| Control 6h | 45.657 | 13.7451 | 0.495637 | 0.159114 |
| ABA 6h | 12.3165 | 3.22702 | 0.142338 | 0.0378992 |
| ACC 6h | 31.7415 | 7.63529 | 0.319741 | 0.0793579 |
| BABA 6h | 101.395 | 16.2827 | 1.09265 | 0.146004 |
| Chitin 6h | 49.2895 | 9.21516 | 0.555728 | 0.0908794 |
| Epi 6h | 54.1564 | 8.5739 | 0.581962 | 0.082876 |
| SA 6h | 21.1514 | 5.83434 | 0.2465 | 0.0489969 |
| Me-JA 6h | 13.4344 | 3.14486 | 0.163677 | 0.0404547 |
Source Transcript PGSC0003DMT400071105 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G78410.1 | +3 | 5e-19 | 79 | 48/113 (42%) | VQ motif-containing protein | chr1:29502728-29503054 FORWARD LENGTH=108 |
