Probe CUST_17492_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17492_PI426222305 | JHI_St_60k_v1 | DMT400068086 | GAGGATACTCCTCCATCTCGATTTACTTGTCAAATACTCGTTTCTTAATTTGATTTCTCC |
All Microarray Probes Designed to Gene DMG400026481
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17492_PI426222305 | JHI_St_60k_v1 | DMT400068086 | GAGGATACTCCTCCATCTCGATTTACTTGTCAAATACTCGTTTCTTAATTTGATTTCTCC |
| CUST_17525_PI426222305 | JHI_St_60k_v1 | DMT400068085 | GATTCGACTGGTCATAAGACGCAGGAGGCAGACTCTAATGGTCATATAGTGGACAATTAA |
Microarray Signals from CUST_17492_PI426222305
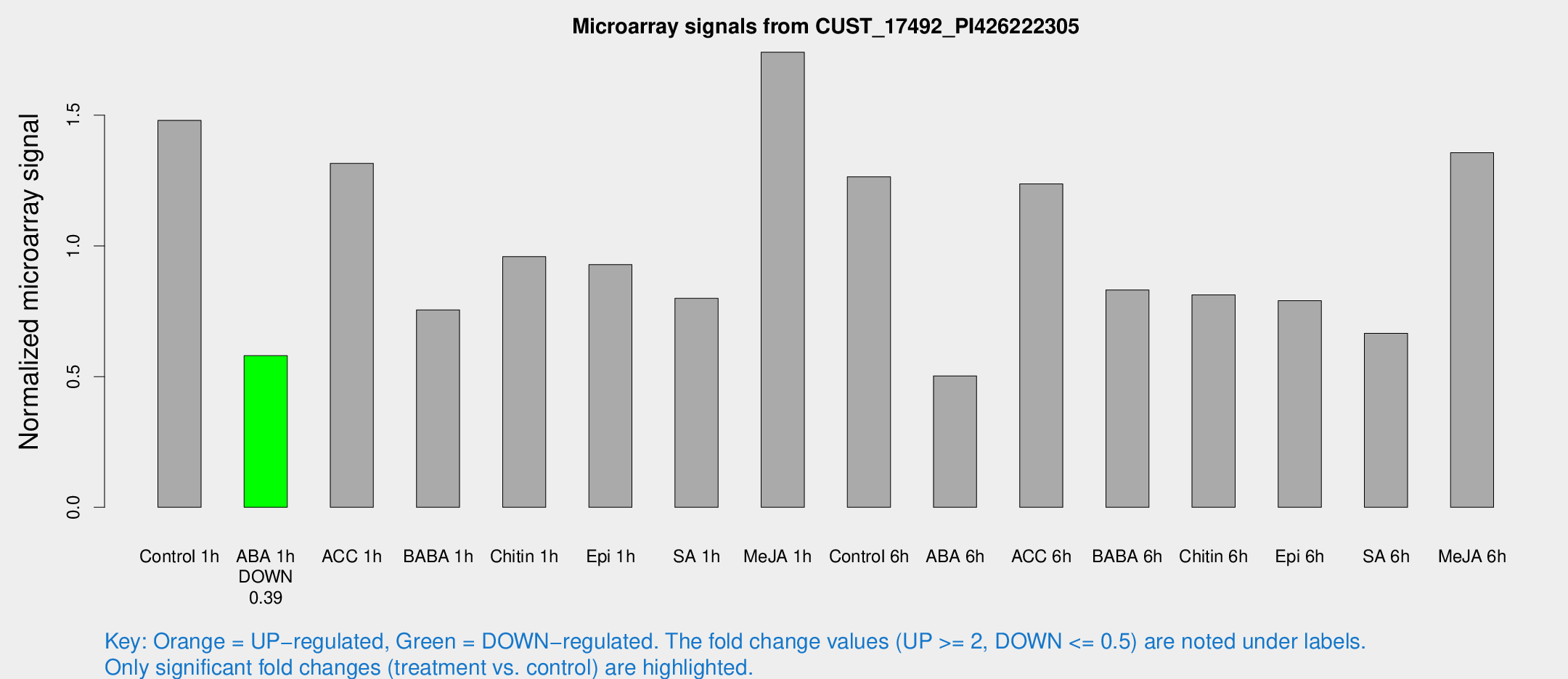
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 255.367 | 31.3128 | 1.48001 | 0.0877041 |
| ABA 1h | 87.3365 | 5.99653 | 0.580126 | 0.0575596 |
| ACC 1h | 232.568 | 27.0817 | 1.31555 | 0.137431 |
| BABA 1h | 129.665 | 27.3081 | 0.755138 | 0.0998229 |
| Chitin 1h | 147.769 | 13.7908 | 0.958864 | 0.127672 |
| Epi 1h | 137.994 | 14.4633 | 0.928225 | 0.107873 |
| SA 1h | 165.723 | 59.0618 | 0.799125 | 0.366156 |
| Me-JA 1h | 242.281 | 14.4494 | 1.74071 | 0.122598 |
| Control 6h | 221.998 | 39.1437 | 1.26445 | 0.162938 |
| ABA 6h | 93.1609 | 13.8131 | 0.502804 | 0.0615209 |
| ACC 6h | 245.387 | 32.1691 | 1.23693 | 0.110744 |
| BABA 6h | 161.835 | 25.2805 | 0.831566 | 0.125058 |
| Chitin 6h | 150.188 | 22.5804 | 0.812851 | 0.0955876 |
| Epi 6h | 152.067 | 9.70646 | 0.790374 | 0.108567 |
| SA 6h | 116.608 | 21.891 | 0.665359 | 0.0621815 |
| Me-JA 6h | 234.074 | 32.0254 | 1.35656 | 0.0812794 |
Source Transcript PGSC0003DMT400068086 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G16760.1 | +3 | 3e-98 | 301 | 157/310 (51%) | Inositol 1,3,4-trisphosphate 5/6-kinase family protein | chr5:5509890-5510849 FORWARD LENGTH=319 |
