Probe CUST_17476_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17476_PI426222305 | JHI_St_60k_v1 | DMT400008327 | GCCCTGTTTTTGCTCTTCTTTAATTCCTCCCAGCTCACGATCTTTGAGTTTGGGTTTTGT |
All Microarray Probes Designed to Gene DMG401003209
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17552_PI426222305 | JHI_St_60k_v1 | DMT400008321 | ATTGATCAACAGATGAACGGTGTTGATTTGAGAAGAAATGGAGGGGTTGTTGATGAGGAT |
| CUST_17476_PI426222305 | JHI_St_60k_v1 | DMT400008327 | GCCCTGTTTTTGCTCTTCTTTAATTCCTCCCAGCTCACGATCTTTGAGTTTGGGTTTTGT |
Microarray Signals from CUST_17476_PI426222305
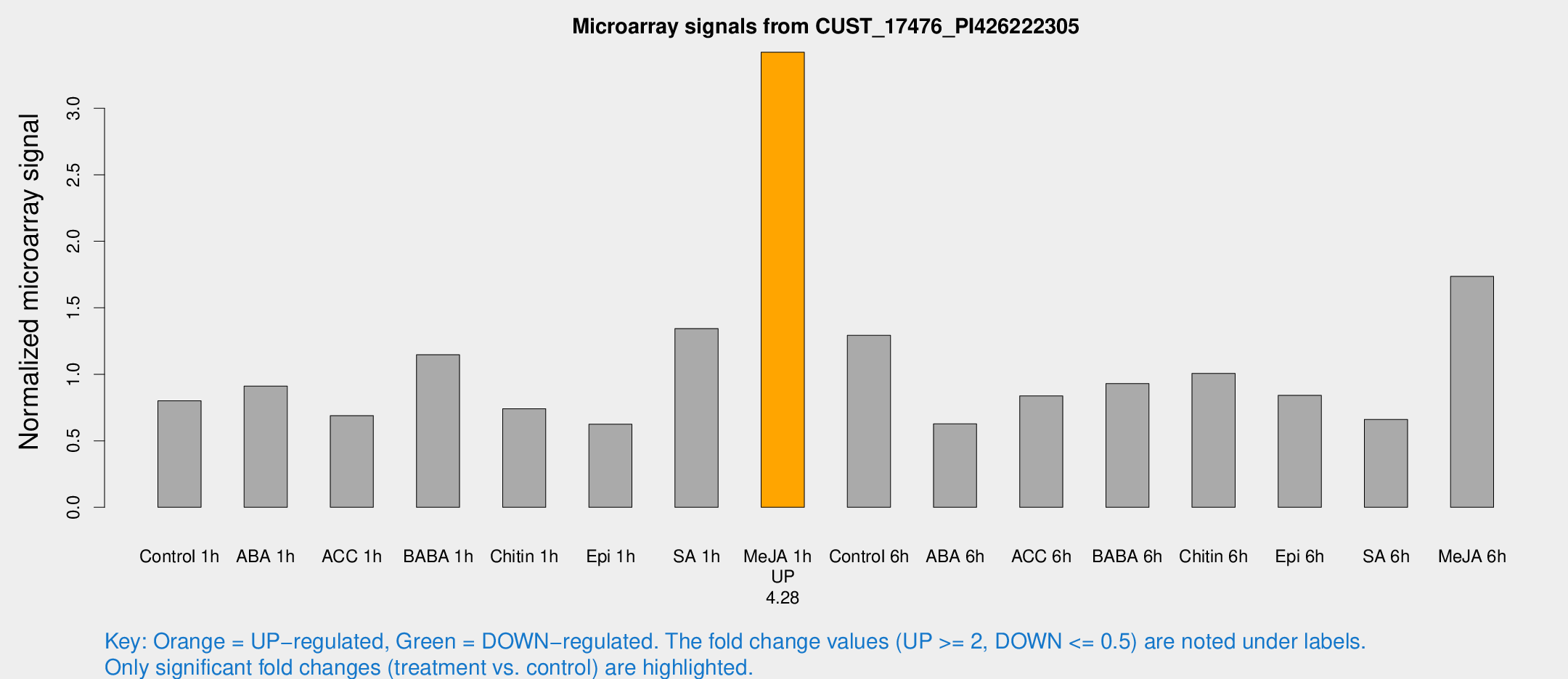
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 36.2946 | 3.98237 | 0.800073 | 0.0884431 |
| ABA 1h | 38.1292 | 7.30835 | 0.911437 | 0.158689 |
| ACC 1h | 35.4253 | 9.9297 | 0.688555 | 0.190478 |
| BABA 1h | 50.2776 | 4.75754 | 1.14698 | 0.109485 |
| Chitin 1h | 30.9372 | 4.64674 | 0.741319 | 0.184221 |
| Epi 1h | 25.8387 | 5.93682 | 0.624551 | 0.166719 |
| SA 1h | 66.0818 | 16.1657 | 1.34354 | 0.31106 |
| Me-JA 1h | 127.218 | 9.53767 | 3.42117 | 0.316629 |
| Control 6h | 62.4358 | 15.7727 | 1.2923 | 0.223855 |
| ABA 6h | 33.2631 | 9.36837 | 0.627387 | 0.196019 |
| ACC 6h | 44.5704 | 7.2866 | 0.836921 | 0.0947729 |
| BABA 6h | 49.8913 | 10.759 | 0.929646 | 0.20609 |
| Chitin 6h | 49.7417 | 7.35206 | 1.00648 | 0.167374 |
| Epi 6h | 48.768 | 17.0825 | 0.841533 | 0.371415 |
| SA 6h | 36.7858 | 14.0037 | 0.660916 | 0.483426 |
| Me-JA 6h | 93.9077 | 32.06 | 1.73538 | 0.767135 |
Source Transcript PGSC0003DMT400008327 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G10610.1 | +2 | 1e-127 | 381 | 197/268 (74%) | CTC-interacting domain 12 | chr4:6557336-6559143 FORWARD LENGTH=336 |
